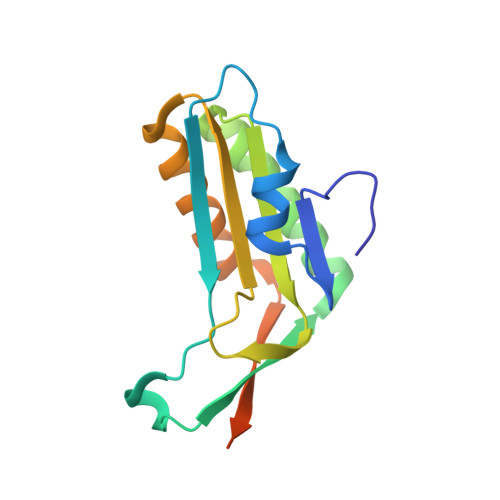
Crystal structure of molybdopterin converting factor subunit 2 (aq_2181) from aquifex aeolicus VF5
Jeyakanthan, J., Kanaujia, S.P., Vasuki Ranjani, C., Sekar, K., Agari, Y., Ebihara, A., Kuramitsu, S., Shinkai, A., Shiro, Y., Yokoyama, S.To be published.