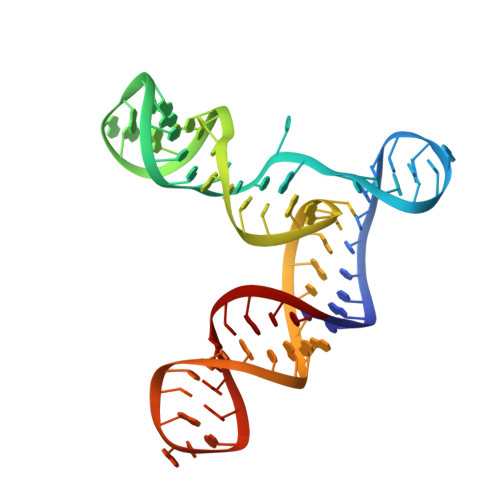The structural basis of ribozyme-catalyzed RNA assembly.
Robertson, M.P., Scott, W.G.(2007) Science 315: 1549-1553
- PubMed: 17363667 Search on PubMed
- DOI: https://doi.org/10.1126/science.1136231
- Primary Citation Related Structures:
2OIU - PubMed Abstract:
Life originated, according to the RNA World hypothesis, from self-replicating ribozymes that catalyzed ligation of RNA fragments. We have solved the 2.6 angstrom crystal structure of a ligase ribozyme that catalyzes regiospecific formation of a 5' to 3' phosphodiester bond between the 5'-triphosphate and the 3'-hydroxyl termini of two RNA fragments. Invariant residues form tertiary contacts that stabilize a flexible stem of the ribozyme at the ligation site, where an essential magnesium ion coordinates three phosphates. The structure of the active site permits us to suggest how transition-state stabilization and a general base may catalyze the ligation reaction required for prebiotic RNA assembly.
- Center for the Molecular Biology of RNA and Department of Chemistry and Biochemistry, Robert L. Sinsheimer Laboratories, University of California, Santa Cruz, Santa Cruz, CA 95064, USA.
Organizational Affiliation: