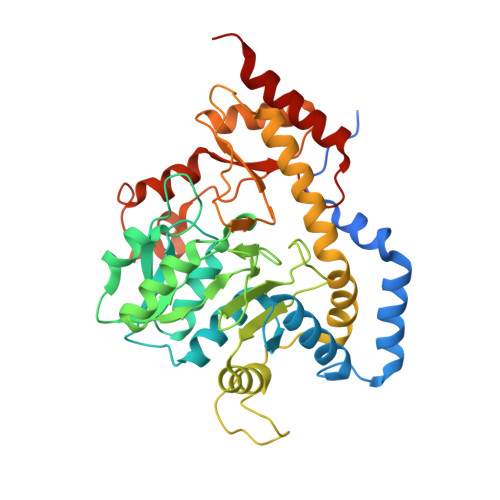
Molecular architecture of DesV from Streptomyces venezuelae: A PLP-dependent transaminase involved in the biosynthesis of the unusual sugar desosamine.
Burgie, E.S., Thoden, J.B., Holden, H.M.(2007) Protein Sci 16: 887-896
- PubMed: 17456741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062711007
- Primary Citation Related Structures:
2OGA, 2OGE - PubMed Abstract:
Desosamine is a 3-(dimethylamino)-3,4,6-trideoxyhexose found in certain macrolide antibiotics such as the commonly prescribed erythromycin. Six enzymes are required for its biosynthesis in Streptomyces venezuelae. The focus of this article is DesV, which catalyzes the PLP-dependent replacement of a 3-keto group with an amino functionality in the fifth step of the pathway. For this study the three-dimensional structures of both the internal aldimine and the ketimine intermediate with glutamate were determined to 2.05 A resolution. DesV is a homodimer with each subunit containing 12 alpha-helical regions and 12 beta-strands that together form three layers of sheet. The structure of the internal aldimine demonstrates that the PLP-cofactor is held in place by residues contributed from both subunits (Asp 164 and Gln 167 from Subunit I and Tyr 221 and Asn 235 from Subunit II). When the ketimine intermediate is present in the active site, the loop defined by Gln 225 to Ser 228 from Subunit II closes down upon the active site. The structure of DesV is similar to another sugar-modifying enzyme referred to as PseC. This enzyme is involved in the biosynthesis of pseudaminic acid, which is a sialic acid-like nonulosonate found in the flagellin of Helicobacter pylori. In the case of PseC, however, the amino group is transferred to the C-4 rather than the C-3 position. Details concerning the structural analysis of DesV and a comparison of its molecular architecture to that of PseC are presented.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: