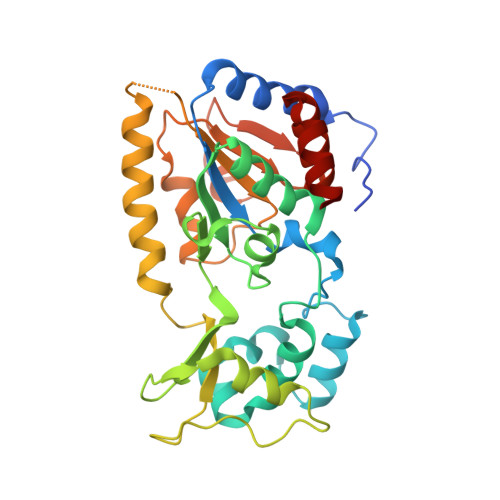
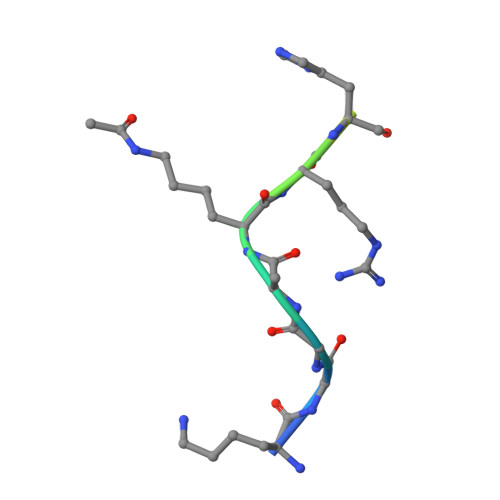
Structural basis for nicotinamide inhibition and base exchange in sir2 enzymes.
Sanders, B.D., Zhao, K., Slama, J.T., Marmorstein, R.(2007) Mol Cell 25: 463-472
- PubMed: 17289592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2006.12.022
- Primary Citation Related Structures:
2OD2, 2OD7, 2OD9, 2QQF, 2QQG - PubMed Abstract:
The Sir2 family of proteins consists of broadly conserved NAD(+)-dependent deacetylases that are implicated in diverse biological processes, including DNA regulation, metabolism, and longevity. Sir2 proteins are regulated in part by the cellular concentrations of a noncompetitive inhibitor, nicotinamide, that reacts with a Sir2 reaction intermediate via a base-exchange reaction to reform NAD(+) at the expense of deacetylation. To gain a mechanistic understanding of nicotinamide inhibition in Sir2 enzymes, we captured the structure of nicotinamide bound to a Sir2 homolog, yeast Hst2, in complex with its acetyl-lysine 16 histone H4 substrate and a reaction intermediate analog, ADP-HPD. Together with related biochemical studies and structures, we identify a nicotinamide inhibition and base-exchange site that is distinct from the so-called "C pocket" binding site for the nicotinamide group of NAD(+). These results provide insights into the Sir2 mechanism of nicotinamide inhibition and have important implications for the development of Sir2-specific effectors.
- The Wistar Institute, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: