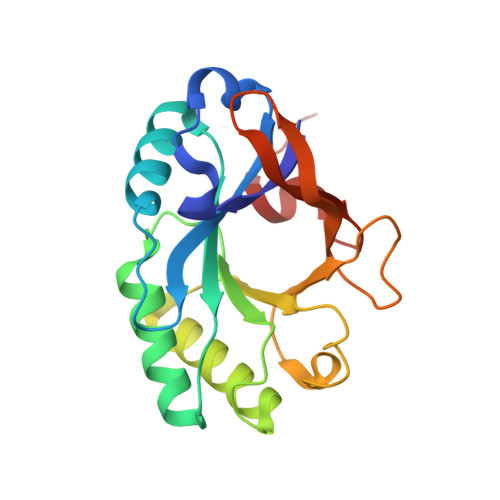
The 1.6 A Crystal Structure of the Catalytic Domain of PlyB, a Bacteriophage Lysin Active Against Bacillus anthracis.
Porter, C.J., Schuch, R., Pelzek, A.J., Buckle, A.M., McGowan, S., Wilce, M.C., Rossjohn, J., Russell, R., Nelson, D., Fischetti, V.A., Whisstock, J.C.(2007) J Mol Biology 366: 540-550
- PubMed: 17182056 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.11.056
- Primary Citation Related Structures:
2NW0 - PubMed Abstract:
Lysins are peptidoglycan hydrolases that are produced by bacteriophage and act to lyse the bacterial host cell wall during progeny phage release. Here, we describe the structure and function of a novel bacteriophage-derived lysin, PlyB, which displays potent lytic activity against the Bacillus anthracis-like strain ATCC 4342. This molecule comprises an N-terminal catalytic domain (PlyB(cat)) and a C-terminal bacterial SH3-like domain, SH3b. It is shown that both domains are required for effective catalytic activity against ATCC 4342. Further, PlyB has specific activity comparable to the phage lysin PlyG, an amidase being developed as a therapeutic against anthrax. In contrast to PlyG, however, the 1.6 A X-ray crystal structure of PlyB(cat) reveals that the catalytic domain adopts the glycosyl hydrolase (GH)-25, rather than phage T7 lysozyme-like fold. PlyB therefore represents a new class of anthrax lysin and a new defensive tool in the armament against anthrax-mediated bioterrorism.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology and The ARC Centre of Excellence for Structural and Functional Microbial Genomics, Monash University, Clayton, Melbourne, VIC 3800, Australia.
Organizational Affiliation: