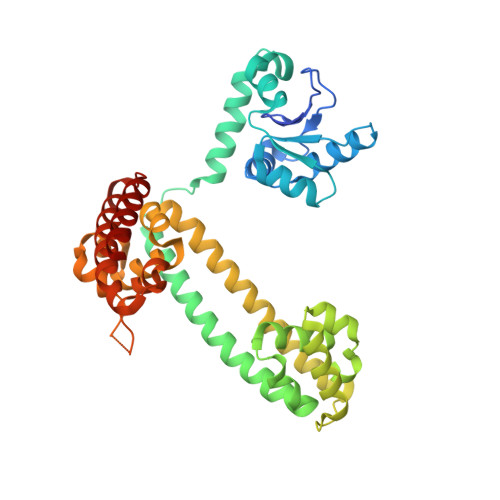
Alternative Conformations of the Archaeal Nop56/58-Fibrillarin Complex Imply Flexibility in Box C/D RNPs.
Oruganti, S., Zhang, Y., Li, H., Robinson, H., Terns, M.P., Terns, R.M., Yang, W., Li, H.(2007) J Mol Biology 371: 1141-1150
- PubMed: 17617422 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.06.029
- Primary Citation Related Structures:
2NNW - PubMed Abstract:
The Nop56/58-fibrillarin heterocomplex is a core protein complex of the box C/D ribonucleoprotein particles that modify and process ribosomal RNAs. The previous crystal structure of the Archaeoglobus fulgidus complex revealed a symmetric dimer of two Nop56/58-fibrillarin complexes linked by the coiled-coil domains of the Nop56/68 proteins. However, because the A. fulgidus Nop56/58 protein lacks some domains found in most other species, it was thought that the bipartite architecture of the heterocomplex was not likely a general phenomenon. Here we report the crystal structure of the Nop56/58-fibrillarin complex bound with methylation cofactor, S-adenosyl-L-methionine from Pyrococcus furiosus, at 2.7 A. The new complex confirms the generality of the previously observed bipartite arrangement. In addition however, the conformation of Nop56/58 in the new structure differs substantially from that in the earlier structure. The distinct conformations of Nop56/58 suggest potential flexibility in Nop56/58. Computational normal mode analysis supports this view. Importantly, fibrillarin is repositioned within the two complexes. We propose that hinge motion within Nop56/58 has important implications for the possibility of simultaneously positioning two catalytic sites at the two target sites of a bipartite box C/D guide RNA.
- Department of Chemistry and Biochemistry, Florida State University, Tallahassee, FL 32306, USA.
Organizational Affiliation: