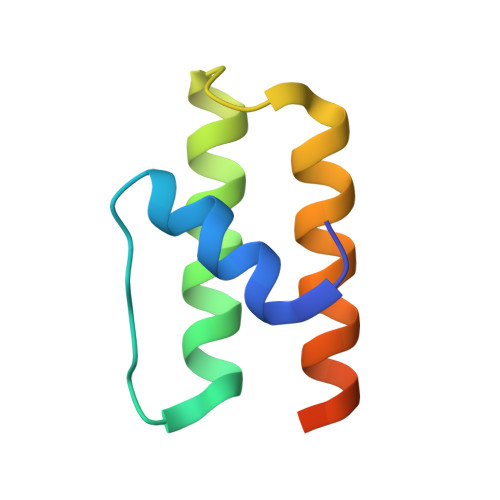
Structure of YqgQ protein from Bacillus subtilis, a conserved hypothetical protein.
Lakshminarasimhan, D., Eswaramoorthy, S., Burley, S.K., Swaminathan, S.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 8-11
- PubMed: 20057058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109047009
- Primary Citation Related Structures:
2NN4 - PubMed Abstract:
The crystal structure of the hypothetical protein YqgQ from Bacillus subtilis has been determined to 2.1 A resolution. The crystals belonged to space group P2(1), with unit-cell parameters a = 51.85, b = 41.25, c = 55.18 A, beta = 113.4 degrees , and contained three protein molecules in the asymmetric unit. The structure was determined by the single-wavelength anomalous dispersion method using selenium-labeled protein and was refined to a final R factor of 24.7% (R(free) = 28.0%). The protein molecule mainly comprises a three-helical bundle. Its putative function is inferred to be single-stranded nucleic acid binding based on sequence and structural homology.
- Biology Department, Brookhaven National Laboratory, Upton, New York 11973, USA.
Organizational Affiliation: