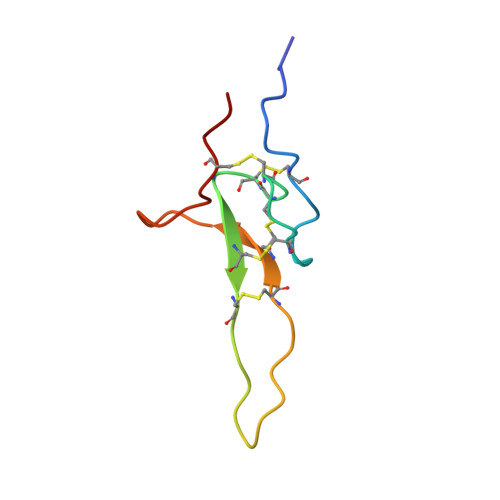
Modular toxin from the lynx spider Oxyopes takobius: Structure of spiderine domains in solution and membrane-mimicking environment.
Nadezhdin, K.D., Romanovskaia, D.D., Sachkova, M.Y., Oparin, P.B., Kovalchuk, S.I., Grishin, E.V., Arseniev, A.S., Vassilevski, A.A.(2017) Protein Sci 26: 611-616
- PubMed: 27997708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3101
- Primary Citation Related Structures:
2N85, 2N86 - PubMed Abstract:
We have recently demonstrated that a common phenomenon in evolution of spider venom composition is the emergence of so-called modular toxins consisting of two domains, each corresponding to a "usual" single-domain toxin. In this article, we describe the structure of two domains that build up a modular toxin named spiderine or OtTx1a from the venom of Oxyopes takobius. Both domains were investigated by solution NMR in water and detergent micelles used to mimic membrane environment. The N-terminal spiderine domain OtTx1a-AMP (41 amino acid residues) contains no cysteines. It is disordered in aqueous solution but in micelles, it assumes a stable amphiphilic structure consisting of two α-helices separated by a flexible linker. On the contrary, the C-terminal domain OtTx1a-ICK (59 residues) is a disulfide-rich polypeptide reticulated by five S-S bridges. It presents a stable structure in water and its core is the inhibitor cystine knot (ICK) or knottin motif that is common among single-domain neurotoxins. OtTx1a-ICK structure is the first knottin with five disulfide bridges and it represents a good reference for the whole oxytoxin family. The affinity of both domains to membranes was measured with NMR using titration by liposome suspensions. In agreement with biological tests, OtTx1a-AMP was found to show high membrane affinity explaining its potent antimicrobial properties.
- M.M. Shemyakin and Yu.A. Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, 16/10 Miklukho-Maklaya, Moscow, 117997, Russia.
Organizational Affiliation: