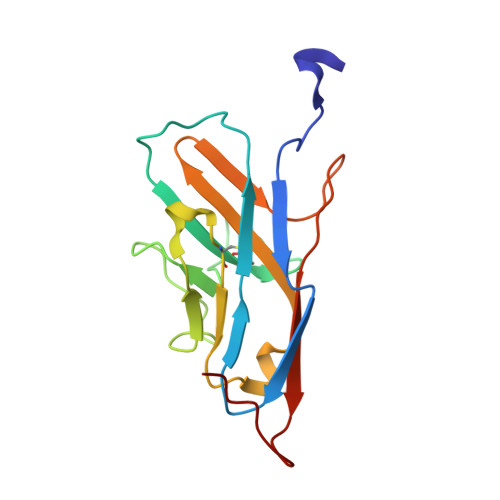
Structural basis for sulfation-dependent self-glycan recognition by the human immune-inhibitory receptor Siglec-8.
Propster, J.M., Yang, F., Rabbani, S., Ernst, B., Allain, F.H., Schubert, M.(2016) Proc Natl Acad Sci U S A 113: E4170-E4179
- PubMed: 27357658 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1602214113
- Primary Citation Related Structures:
2N7A, 2N7B - PubMed Abstract:
Siglec-8 is a human immune-inhibitory receptor that, when engaged by specific self-glycans, triggers eosinophil apoptosis and inhibits mast cell degranulation, providing an endogenous mechanism to down-regulate immune responses of these central inflammatory effector cells. Here we used solution NMR spectroscopy to dissect the fine specificity of Siglec-8 toward different sialylated and sulfated carbohydrate ligands and determined the structure of the Siglec-8 lectin domain in complex with its prime glycan target 6'-sulfo sialyl Lewis(x) A canonical motif for sialic acid recognition, extended by a secondary motif formed by unique loop regions, recognizing 6-O-sulfated galactose dictates tight specificity distinct from other Siglec family members and any other endogenous glycan recognition receptors. Structure-guided mutagenesis revealed key contacts of both interfaces to be equally essential for binding. Our work provides critical structural and mechanistic insights into how Siglec-8 selectively recognizes its glycan target, rationalizes the functional impact of site-specific glycan sulfation in modulating this lectin-glycan interaction, and will enable the rational design of Siglec-8-targeted agonists to treat eosinophil- and mast cell-related allergic and inflammatory diseases, such as asthma.
- Institute of Molecular Biology and Biophysics, ETH Zurich, 8093 Zurich, Switzerland;
Organizational Affiliation: