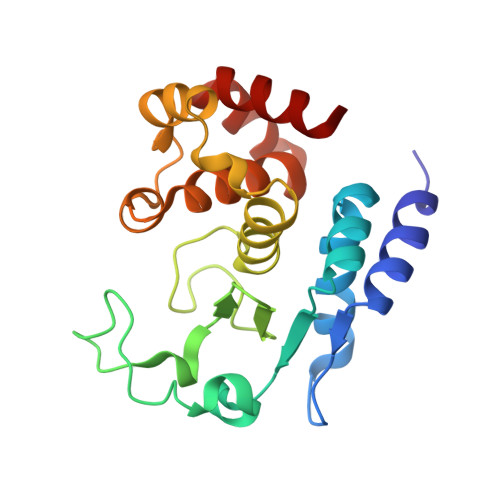
Structural characterization of zinc-bound Zmp1, a zinc-dependent metalloprotease secreted by Clostridium difficile.
Rubino, J.T., Martinelli, M., Cantini, F., Castagnetti, A., Leuzzi, R., Banci, L., Scarselli, M.(2016) J Biol Inorg Chem 21: 185-196
- PubMed: 26711661 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-015-1319-6
- Primary Citation Related Structures:
2N6J - PubMed Abstract:
Proteases are commonly secreted by microorganisms. In some pathogens, they can play a series of functional roles during infection, including maturation of cell surface or extracellular virulence factors, interference with host cell signaling, massive host tissue destruction, and dissolution of infection-limiting clots through degradation of the host proteins devoted to the coagulation cascade. We previously reported the identification and characterization of Zmp1, a zinc-dependent metalloprotease secreted by Clostridium difficile, demonstrated that Zmp1 is able to degrade fibrinogen in vitro, and identified two residues necessary to the catalytic activity. In the present work, we solved the solution structure of Zmp1 by Nuclear Magnetic Resonance (NMR) and compared it with the recently solved X-ray structures of substrate-bound and substrate-free Zmp1, highlighting similarities and differences. We also combined the structural characterization to biochemical assays and site-directed mutagenesis, to provide new insights into the catalytic site and on the residues responsible for substrate specificity. The Zmp1 structure showed similarity to the catalytic domain of Anthrax Lethal Factor of Bacillus anthracis. Analogies and differences in the catalytic and in the substrate-binding sites of the two proteins are discussed.
- Magnetic Resonance Center, University of Florence, Via L. Sacconi 6, 50019, Sesto Fiorentino, Italy.
Organizational Affiliation: