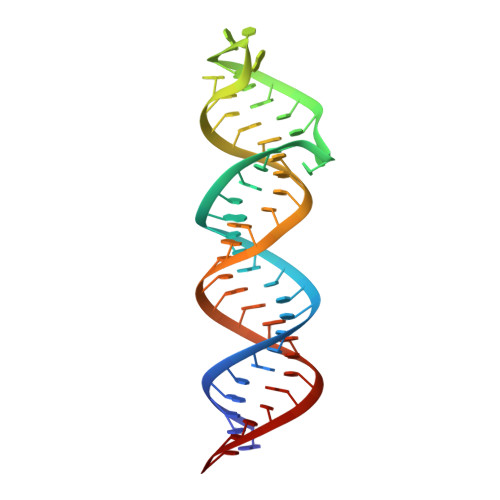
Solution Structure of the HIV-1 Intron Splicing Silencer and Its Interactions with the UP1 Domain of Heterogeneous Nuclear Ribonucleoprotein (hnRNP) A1.
Jain, N., Morgan, C.E., Rife, B.D., Salemi, M., Tolbert, B.S.(2016) J Biological Chem 291: 2331-2344
- PubMed: 26607354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.674564
- Primary Citation Related Structures:
2N4L - PubMed Abstract:
Splicing patterns in human immunodeficiency virus type 1 (HIV-1) are maintained through cis regulatory elements that recruit antagonistic host RNA-binding proteins. The activity of the 3' acceptor site A7 is tightly regulated through a complex network of an intronic splicing silencer (ISS), a bipartite exonic splicing silencer (ESS3a/b), and an exonic splicing enhancer (ESE3). Because HIV-1 splicing depends on protein-RNA interactions, it is important to know the tertiary structures surrounding the splice sites. Herein, we present the NMR solution structure of the phylogenetically conserved ISS stem loop. ISS adopts a stable structure consisting of conserved UG wobble pairs, a folded 2X2 (GU/UA) internal loop, a UU bulge, and a flexible AGUGA apical loop. Calorimetric and biochemical titrations indicate that the UP1 domain of heterogeneous nuclear ribonucleoprotein A1 binds the ISS apical loop site-specifically and with nanomolar affinity. Collectively, this work provides additional insights into how HIV-1 uses a conserved RNA structure to commandeer a host RNA-binding protein.
- From the Department of Chemistry, Case Western Reserve University, Cleveland, Ohio 44106-7078 and.
Organizational Affiliation: