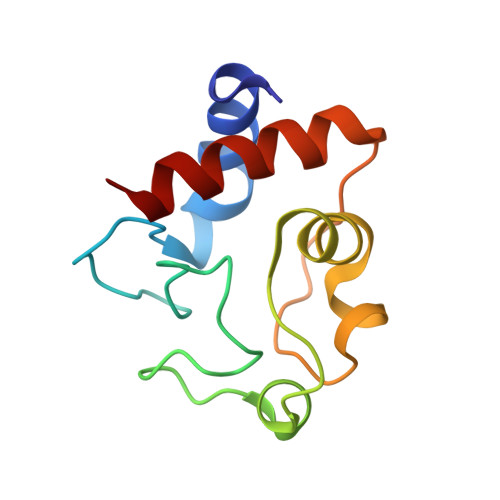
Defining the Apoptotic Trigger: THE INTERACTION OF CYTOCHROME c AND CARDIOLIPIN.
O'Brien, E.S., Nucci, N.V., Fuglestad, B., Tommos, C., Wand, A.J.(2015) J Biological Chem 290: 30879-30887
- PubMed: 26487716 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.689406
- Primary Citation Related Structures:
2N3B - PubMed Abstract:
The interaction between cytochrome c and the anionic lipid cardiolipin has been proposed as a primary event in the apoptotic signaling cascade. Numerous studies that have examined the interaction of cytochrome c with cardiolipin embedded in a variety of model phospholipid membranes have suggested that partial unfolding of the protein is a precursor to the apoptotic response. However, these studies lacked site resolution and used model systems with negligible or a positive membrane curvature, which is distinct from the large negative curvature of the invaginations of the inner mitochondrial membrane where cytochrome c resides. We have used reverse micelle encapsulation to mimic the potential effects of confinement on the interaction of cytochrome c with cardiolipin. Encapsulation of oxidized horse cytochrome c in 1-decanoyl-rac-glycerol/lauryldimethylamine-N-oxide/hexanol reverse micelles prepared in pentane yields NMR spectra essentially identical to the protein in free aqueous solution. The structure of encapsulated ferricytochrome c was determined to high precision (
bb ∼ 0.23 Å) using NMR-based methods and is closely similar to the cryogenic crystal structure ( bb ∼ 1.2 Å). Incorporation of cardiolipin into the reverse micelle surfactant shell causes localized chemical shift perturbations of the encapsulated protein, providing the first view of the cardiolipin/cytochrome c interaction interface at atomic resolution. Three distinct sites of interaction are detected: the so-called A- and L-sites, plus a previously undocumented interaction centered on residues Phe-36, Gly-37, Thr-58, Trp-59, and Lys-60. Importantly, in distinct contrast to earlier studies of this interaction, the protein is not significantly disturbed by the binding of cardiolipin in the context of the reverse micelle. - From the Johnson Research Foundation and Department of Biochemistry and Biophysics, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6059.
Organizational Affiliation: