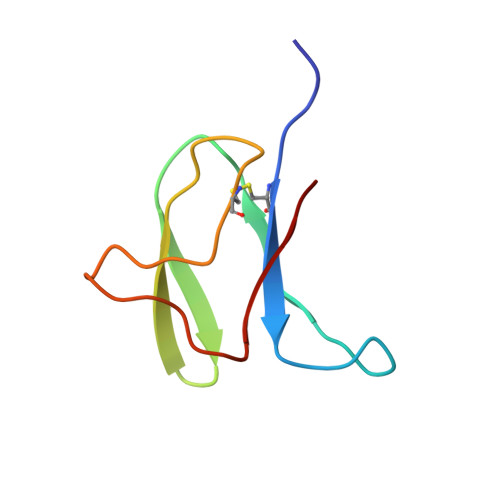
Solution structure of an avirulence protein, AVR-Pia, from Magnaporthe oryzae
Ose, T., Oikawa, A., Nakamura, Y., Maenaka, K., Higuchi, Y., Satoh, Y., Fujiwara, S., Demura, M., Sone, T., Kamiya, M.(2015) J Biomol NMR 63: 229-235
- PubMed: 26362280 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-015-9979-7
- Primary Citation Related Structures:
2N37 - Faculty of Pharmaceutical Sciences, Hokkaido University, N12, W6, Kita-ku, Sapporo, Hokkaido, 060-0812, Japan.
Organizational Affiliation: