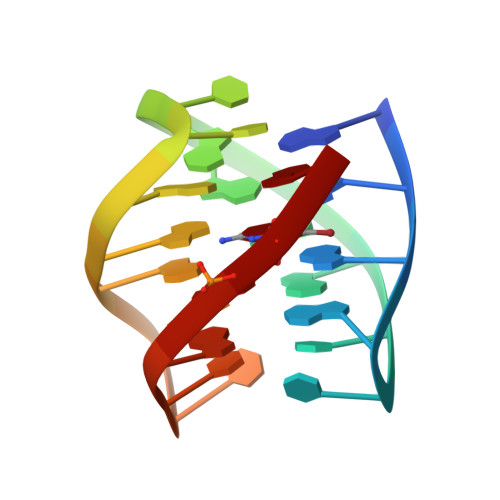
Solution structure of a DNA quadruplex containing ALS and FTD related GGGGCC repeat stabilized by 8-bromodeoxyguanosine substitution.
Brcic, J., Plavec, J.(2015) Nucleic Acids Res 43: 8590-8600
- PubMed: 26253741
- DOI: https://doi.org/10.1093/nar/gkv815
- Primary Citation Related Structures:
2N2D - PubMed Abstract:
A prolonged expansion of GGGGCC repeat within non-coding region of C9orf72 gene has been identified as the most common cause of familial amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD), which are devastating neurodegenerative disorders. Formation of unusual secondary structures within expanded GGGGCC repeat, including DNA and RNA G-quadruplexes and R-loops was proposed to drive ALS and FTD pathogenesis. Initial NMR investigation on DNA oligonucleotides with four repeat units as the shortest model with the ability to form an unimolecular G-quadruplex indicated their folding into multiple G-quadruplex structures in the presence of K(+) ions. Single dG to 8Br-dG substitution at position 21 in oligonucleotide d[(G4C2)3G4] and careful optimization of folding conditions enabled formation of mostly a single G-quadruplex species, which enabled determination of a high-resolution structure with NMR. G-quadruplex structure adopted by d[(G4C2)3GG(Br)GG] is composed of four G-quartets, which are connected by three edgewise C-C loops. All four strands adopt antiparallel orientation to one another and have alternating syn-anti progression of glycosidic conformation of guanine residues. One of the cytosines in every loop is stacked upon the G-quartet contributing to a very compact and stable structure.
- Slovenian NMR Center, National Institute of Chemistry, Hajdrihova 19, SI-1000, Ljubljana, Slovenia.
Organizational Affiliation: