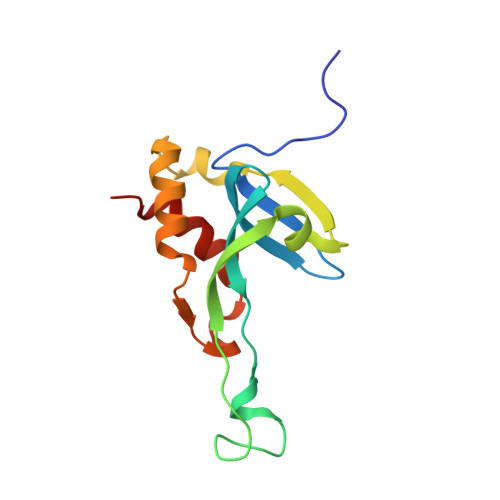
Structural and functional characterization of tumor suppressors TIG3 and H-REV107.
Wei, H., Wang, L., Ren, X., Yu, W., Lin, J., Jin, C., Xia, B.(2015) FEBS Lett 589: 1179-1186
- PubMed: 25871522 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2015.04.002
- Primary Citation Related Structures:
2MY9 - PubMed Abstract:
H-REV107-like family proteins TIG3 and H-REV107 are class II tumor suppressors. Here we report that the C-terminal domains (CTDs) of TIG3 and H-REV107 can induce HeLa cell death independently. The N-terminal domain (NTD) of TIG3 enhances the cell death inducing ability of CTD, while NTD of H-REV107 plays an inhibitory role. The solution structure of TIG3 NTD is very similar to that of H-REV107 in overall fold. However, the CTD binding regions on NTD are different between TIG3 and H-REV107, which may explain their functional difference. As a result, the flexible main loop of H-REV107, but not that of TIG3, is critical for its NTD to modulate its CTD in inducing cell death.
- Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing 100871, China; School of Life Sciences, Peking University, Beijing 100871, China.
Organizational Affiliation: