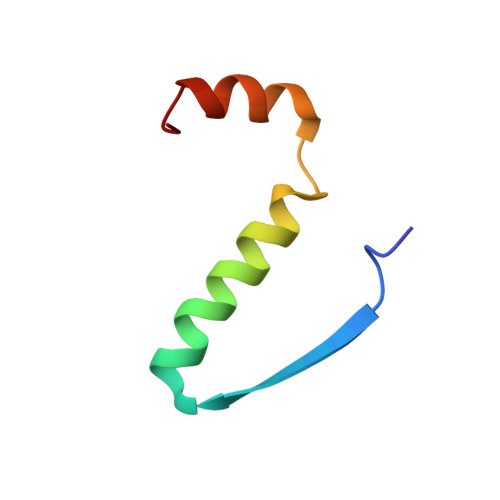
Structural investigations of the p53/p73 homologs from the tunicate species Ciona intestinalis reveal the sequence requirements for the formation of a tetramerization domain.
Heering, J., Jonker, H.R., Lohr, F., Schwalbe, H., Dotsch, V.(2016) Protein Sci 25: 410-422
- PubMed: 26473758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2830
- Primary Citation Related Structures:
2MW4 - PubMed Abstract:
Most members of the p53 family of transcription factors form tetramers. Responsible for determining the oligomeric state is a short oligomerization domain consisting of one β-strand and one α-helix. With the exception of human p53 all other family members investigated so far contain a second α-helix as part of their tetramerization domain. Here we have used nuclear magnetic resonance spectroscopy to characterize the oligomerization domains of the two p53-like proteins from the tunicate Ciona intestinalis, representing the closest living relative of vertebrates. Structure determination reveals for one of the two proteins a new type of packing of this second α-helix on the core domain that was not predicted based on the sequence, while the other protein does not form a second helix despite the presence of crucial residues that are conserved in all other family members that form a second helix. By mutational analysis, we identify a proline as well as large hydrophobic residues in the hinge region between both helices as the crucial determinant for the formation of a second helix.
- Institute of Biophysical Chemistry and Center for Biomolecular Magnetic Resonance, Goethe University, D-60438 Frankfurt/Main, Germany.
Organizational Affiliation: