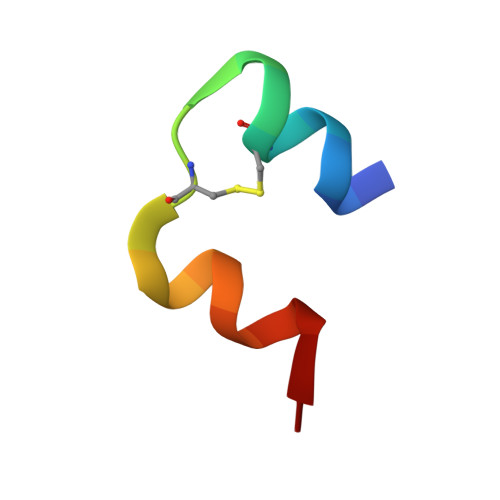
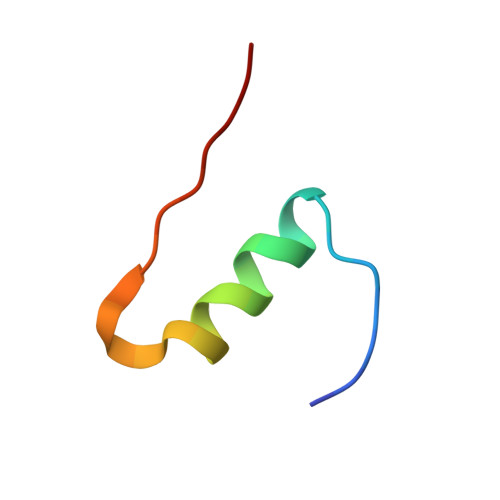
Structural and Functional Study of the GlnB22-Insulin Mutant Responsible for Maturity-Onset Diabetes of the Young.
Krizkova, K., Veverka, V., Maletinska, L., Hexnerova, R., Brzozowski, A.M., Jiracek, J., Zakova, L.(2014) PLoS One 9: e112883-e112883
- PubMed: 25423173 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0112883
- Primary Citation Related Structures:
2MVC, 2MVD - PubMed Abstract:
The insulin gene mutation c.137G>A (R46Q), which changes an arginine at the B22 position of the mature hormone to glutamine, causes the monogenic diabetes variant maturity-onset diabetes of the young (MODY). In MODY patients, this mutation is heterozygous, and both mutant and wild-type (WT) human insulin are produced simultaneously. However, the patients often depend on administration of exogenous insulin. In this study, we chemically synthesized the MODY mutant [GlnB22]-insulin and characterized its biological and structural properties. The chemical synthesis of this insulin analogue revealed that its folding ability is severely impaired. In vitro and in vivo tests showed that its binding affinity and biological activity are reduced (both approximately 20% that of human insulin). Comparison of the solution structure of [GlnB22]-insulin with the solution structure of native human insulin revealed that the most significant structural effect of the mutation is distortion of the B20-B23 β-turn, leading to liberation of the B chain C-terminus from the protein core. The distortion of the B20-B23 β-turn is caused by the extended conformational freedom of the GlnB22 side chain, which is no longer anchored in a hydrogen bonding network like the native ArgB22. The partially disordered [GlnB22]-insulin structure appears to be one reason for the reduced binding potency of this mutant and may also be responsible for its low folding efficiency in vivo. The altered orientation and flexibility of the B20-B23 β-turn may interfere with the formation of disulfide bonds in proinsulin bearing the R46Q (GlnB22) mutation. This may also have a negative effect on the WT proinsulin simultaneously biosynthesized in β-cells and therefore play a major role in the development of MODY in patients producing [GlnB22]-insulin.
- Institute of Organic Chemistry and Biochemistry, Academy of Sciences of the Czech Republic, v.v.i., Flemingovo nám. 2, 166 10 Prague 6, Czech Republic.
Organizational Affiliation: