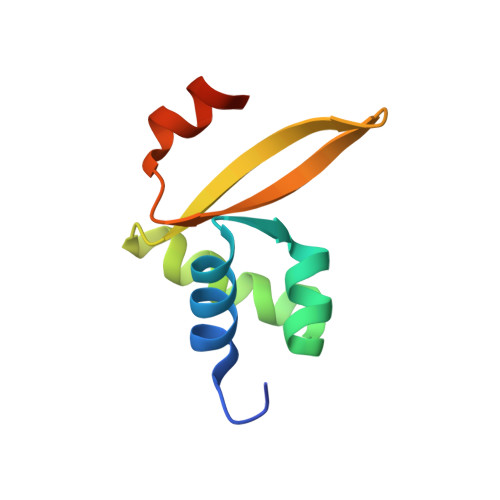
Structural and functional studies of Stf76 from the Sulfolobus islandicus plasmid-virus pSSVx: a novel peculiar member of the winged helix-turn-helix transcription factor family.
Contursi, P., Farina, B., Pirone, L., Fusco, S., Russo, L., Bartolucci, S., Fattorusso, R., Pedone, E.(2014) Nucleic Acids Res 42: 5993-6011
- PubMed: 24682827 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku215
- Primary Citation Related Structures:
2MLG - PubMed Abstract:
The hybrid plasmid-virus pSSVx from Sulfolobus islandicus presents an open reading frame encoding a 76 amino acid protein, namely Stf76, that does not show significant sequence homology with any protein with known 3D structure. The recombinant protein recognizes specifically two DNA-binding sites located in its own promoter, thus suggesting an auto-regulated role of its expression. Circular dichroism, spectrofluorimetric, light scattering and isothermal titration calorimetry experiments indicated a 2:1 molar ratio (protein:DNA) upon binding to the DNA target containing a single site. Furthermore, the solution structure of Stf76, determined by nuclear magnetic resonance (NMR) using chemical shift Rosetta software, has shown that the protein assumes a winged helix-turn-helix fold. NMR chemical shift perturbation analysis has been performed for the identification of the residues responsible for DNA interaction. In addition, a model of the Stf76-DNA complex has been built using as template a structurally related homolog.
- Dipartimento di Biologia, Università degli Studi di Napoli "Federico II", Napoli 80126, Italy.
Organizational Affiliation: