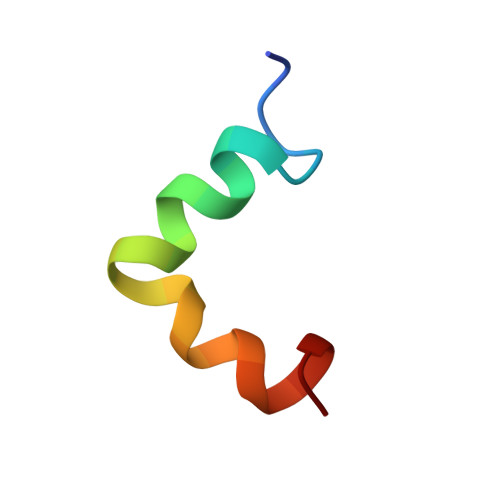
A kinked antimicrobial peptide from Bombina maxima. I. Three-dimensional structure determined by NMR in membrane-mimicking environments.
Toke, O., Banoczi, Z., Kiraly, P., Heinzmann, R., Burck, J., Ulrich, A.S., Hudecz, F.(2011) Eur Biophys J 40: 447-462
- PubMed: 21234559 Search on PubMed
- DOI: https://doi.org/10.1007/s00249-010-0657-0
- Primary Citation Related Structures:
2MHW - PubMed Abstract:
Maximin-4 is a 27-residue cationic antimicrobial peptide exhibiting selectivity for bacterial cells. As part of the innate defense system in the Chinese red-belly toad, its mode of action is thought to be ion channel or pore formation and dissipation of the electrochemical gradient across the pathogenic cell membrane. Here we present the high-resolution structure of maximin-4 in two different membrane mimetics, sodium dodecyl sulfate micelles and 50% methanol, as determined by (1)H solution NMR spectroscopy. In both environments, the peptide chain adopts a helix-break-helix conformation following a highly disordered N-terminal segment. Despite the similarities in the overall topology of the two structures, major differences are observed in terms of the interactions stabilizing the kink region and the arrangement of the four lysine residues. This has a marked influence on the shape and charge distribution of the molecule and may have implications for the bacterial selectivity of the peptide. The solution NMR results are complemented by CD spectroscopy and solid-state NMR experiments in lipid bilayers, both confirming the predominantly helical conformation of the peptide. As a first step in elucidating the membrane interactions of maximin-4, our study contributes to a better understanding of the mode of action of antimicrobial peptides and the factors governing their selectivity.
- Institute of Structural Chemistry, Chemical Research Center of the Hungarian Academy of Sciences, 59-67 Pusztaszeri út, 1025 Budapest, Hungary. toke@chemres.hu
Organizational Affiliation: