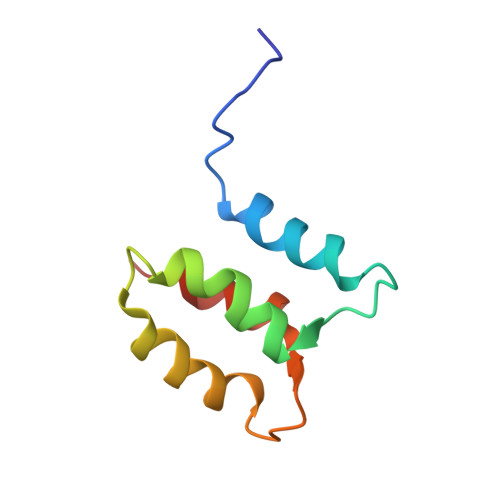
Structure of the Single-lobe Myosin Light Chain C in Complex with the Light Chain-binding Domains of Myosin-1C Provides Insights into Divergent IQ Motif Recognition.
Langelaan, D.N., Liburd, J., Yang, Y., Miller, E., Chitayat, S., Crawley, S.W., Cote, G.P., Smith, S.P.(2016) J Biological Chem 291: 19607-19617
- PubMed: 27466369 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.746313
- Primary Citation Related Structures:
2M8U, 5IBW - PubMed Abstract:
Myosin light chains are key regulators of class 1 myosins and typically comprise two domains, with calmodulin being the archetypal example. They bind IQ motifs within the myosin neck region and amplify conformational changes in the motor domain. A single lobe light chain, myosin light chain C (MlcC), was recently identified and shown to specifically bind to two sequentially divergent IQ motifs of the Dictyostelium myosin-1C. To provide a molecular basis of this interaction, the structures of apo-MlcC and a 2:1 MlcC·myosin-1C neck complex were determined. The two non-functional EF-hand motifs of MlcC pack together to form a globular four-helix bundle that opens up to expose a central hydrophobic groove, which interacts with the N-terminal portion of the divergent IQ1 and IQ2 motifs. The N- and C-terminal regions of MlcC make critical contacts that contribute to its specific interactions with the myosin-1C divergent IQ motifs, which are contacts that deviate from the traditional mode of calmodulin-IQ recognition.
- From the Department of Biomedical and Molecular Sciences, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: