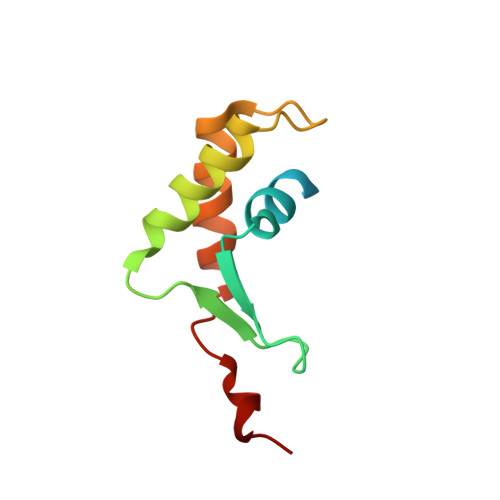
The Solution Structures of Two Prophage Homologues of the Bacteriophage lambda Ea8.5 Protein Reveal a Newly Discovered Hybrid Homeodomain/Zinc-Finger Fold.
Kwan, J.J., Smirnova, E., Khazai, S., Evanics, F., Maxwell, K.L., Donaldson, L.W.(2013) Biochemistry 52: 3612-3614
- PubMed: 23672713 Search on PubMed
- DOI: https://doi.org/10.1021/bi400543w
- Primary Citation Related Structures:
2M7A, 2M7B - PubMed Abstract:
A cluster of genes in the exoxis region of bacteriophage λ are capable of inhibiting the initiation of DNA synthesis in Escherichia coli. The most indispensible gene in this region is ea8.5. Here, we report the nuclear magnetic resonance structures of two ea8.5 orthologs from enteropathogenic E. coli and Pseudomonas putida prophages. Both proteins are characterized by a fused homeodomain/zinc-finger fold that escaped detection by primary sequence search methods. While these folds are both associated with a nucleic acid binding function, the amino acid composition suggests otherwise, leading to the possibility that Ea8.5 associates with other viral and host proteins.
- Department of Biology, York University , 4700 Keele Street, Toronto, ON M3J1P3, Canada.
Organizational Affiliation: