Analysis of the RelA:CBP/p300 Interaction Reveals Its Involvement in NF-kappa B-Driven Transcription.
Mukherjee, S.P., Behar, M., Birnbaum, H.A., Hoffmann, A., Wright, P.E., Ghosh, G.(2013) PLoS Biol 11: e1001647-e1001647
- PubMed: 24019758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1001647
- Primary Citation Related Structures:
2LWW - PubMed Abstract:
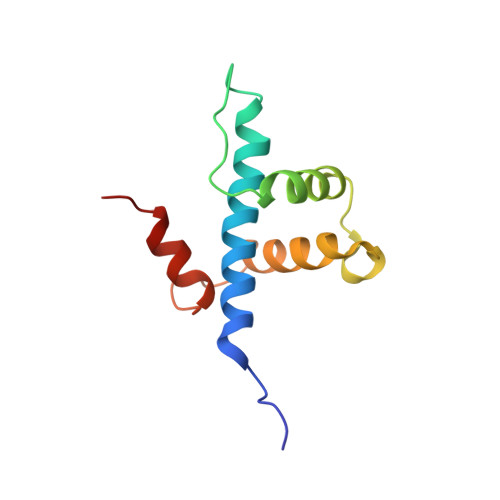
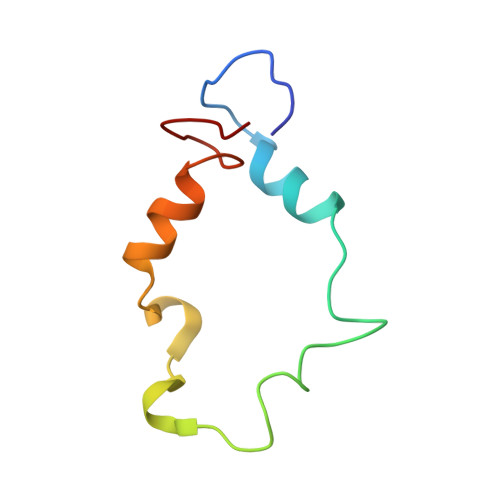
NF-κB plays a vital role in cellular immune and inflammatory response, survival, and proliferation by regulating the transcription of various genes involved in these processes. To activate transcription, RelA (a prominent NF-κB family member) interacts with transcriptional co-activators like CREB-binding protein (CBP) and its paralog p300 in addition to its cognate κB sites on the promoter/enhancer regions of DNA. The RelA:CBP/p300 complex is comprised of two components--first, DNA binding domain of RelA interacts with the KIX domain of CBP/p300, and second, the transcriptional activation domain (TAD) of RelA binds to the TAZ1 domain of CBP/p300. A phosphorylation event of a well-conserved RelA(Ser276) is prerequisite for the former interaction to occur and is considered a decisive factor for the overall RelA:CBP/p300 interaction. The role of the latter interaction in the transcription of RelA-activated genes remains unclear. Here we provide the solution structure of the latter component of the RelA:CBP complex by NMR spectroscopy. The structure reveals the folding of RelA-TA2 (a section of TAD) upon binding to TAZ1 through its well-conserved hydrophobic sites in a series of grooves on the TAZ1 surface. The structural analysis coupled with the mechanistic studies by mutational and isothermal calorimetric analyses allowed the design of RelA-mutants that selectively abrogated the two distinct components of the RelA:CBP/p300 interaction. Detailed studies of these RelA mutants using cell-based techniques, mathematical modeling, and genome-wide gene expression analysis showed that a major set of the RelA-activated genes, larger than previously believed, is affected by this interaction. We further show how the RelA:CBP/p300 interaction controls the nuclear response of NF-κB through the negative feedback loop of NF-κB pathway. Additionally, chromatin analyses of RelA target gene promoters showed constitutive recruitment of CBP/p300, thus indicating a possible role of CBP/p300 in recruitment of RelA to its target promoter sites.
- Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, California, United States of America ; Department of Molecular Biology and The Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, California, United States of America.
Organizational Affiliation: