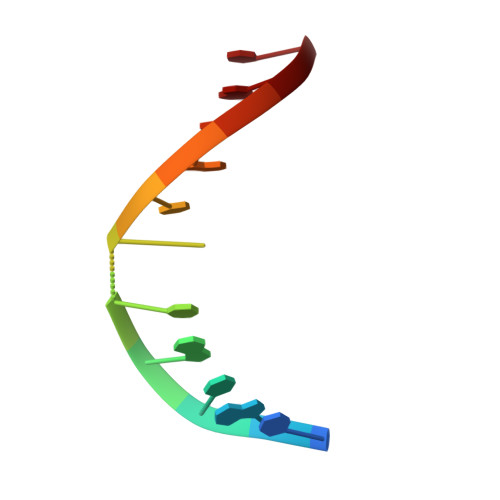
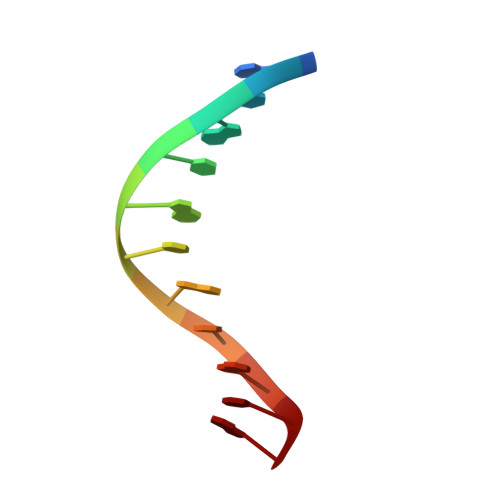
Solution Structure of Duplex DNA Containing a b-Carba-Fapy-dG Lesion
Lukin, M., Zaliznyak, T., Attaluri, S., Johnson, F., de Los Santos, C.(2012) Chem Res Toxicol 25: 2423-2431
- PubMed: 22897814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/tx300290b
- Primary Citation Related Structures:
2LWM, 2LWN, 2LWO - PubMed Abstract:
The addition of hydroxyl radicals to the C8 position of guanine can lead to the formation of a 2,6-diamino-4-hydroxy-5-formamido-2'-deoxypyrimidine (Fapy-dG) lesion, whose endogenous levels in cellular DNA rival those of 8-oxo-7,8-dihydroxy-2'-deoxyguanosine. Despite its prevalence, the structure of duplex DNA containing Fapy-dG is unknown. We have prepared an undecameric duplex containing a centrally located β-cFapy-dG residue paired to dC and determined its solution structure by high-resolution NMR spectroscopy and restrained molecular dynamic simulations. The damaged duplex adopts a right-handed helical structure with all residues in an anti conformation, forming Watson-Crick base pair alignments, and 2-deoxyribose conformations in the C2'-endo/C1'-exo range. The formamido group of Fapy rotates out of the pyrimidine plane and is present in the Z and E configurations that equilibrate with an approximate 2:1 population ratio. The two isomeric duplexes show similar lesion-induced deviations from a canonical B-from DNA conformation that are minor and limited to the central three-base-pair segment of the duplex, affecting the stacking interactions with the 5-lesion-neighboring residue. We discuss the implications of our observations for translesion synthesis during DNA replication and the recognition of Fapy-dG by DNA glycosylases.
- Department of Pharmacological Sciences, Stony Brook University-School of Medicine, Stony Brook, NY 11794-8651, USA.
Organizational Affiliation: