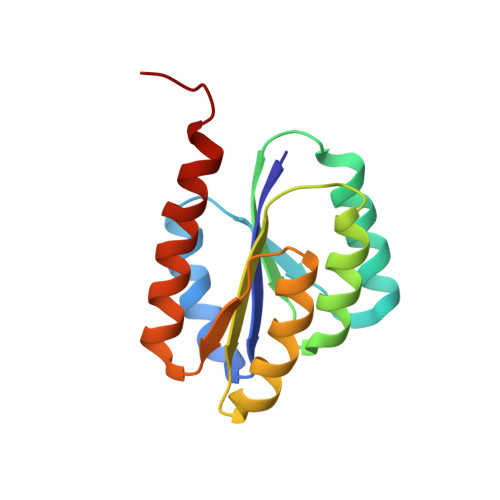
Solution NMR structure of de novo designed protein, p-loop ntpase fold, northeast structural genomics consortium target or136
Liu, G., Koga, N., Koga, R., Xiao, R., Lee, H., Janjua, H., Kohan, E., Acton, T.B., Everett, J.K., Baker, D., Montelione, G.T.To be published.