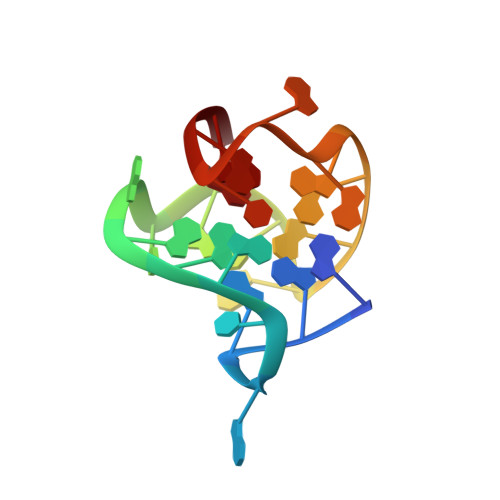
Solution-state structure of an intramolecular G-quadruplex with propeller, diagonal and edgewise loops.
Marusic, M., Sket, P., Bauer, L., Viglasky, V., Plavec, J.(2012) Nucleic Acids Res 40: 6946-6956
- PubMed: 22532609 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks329
- Primary Citation Related Structures:
2LOD - PubMed Abstract:
We herein report on the formation and high-resolution NMR solution-state structure determination of a G-quadruplex adopted by d[G(3)ATG(3)ACACAG(4)ACG(3)] comprised of four G-tracts with the third one consisting of four guanines that are intervened with non-G streches of different lengths. A single intramolecular antiparallel (3+1) G-quadruplex exhibits three stacked G-quartets connected with propeller, diagonal and edgewise loops of different lengths. The propeller and edgewise loops are well structured, whereas the longer diagonal loop is more flexible. To the best of our knowledge, this is the first high-resolution G-quadruplex structure where all of the three main loop types are present.
- Slovenian NMR Center, National Institute of Chemistry, Hajdrihova 19, Slovenia.
Organizational Affiliation: