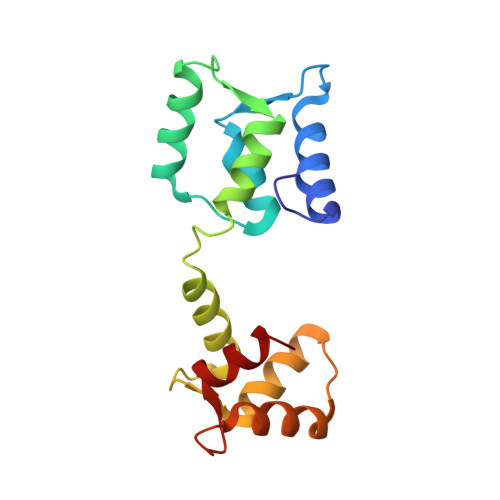
Structure of androcam supports specialized interactions with myosin VI.
Joshi, M.K., Moran, S., Beckingham, K.M., Mackenzie, K.R.(2012) Proc Natl Acad Sci U S A 109: 13290-13295
- PubMed: 22851764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1209730109
- Primary Citation Related Structures:
2LMT, 2LMU, 2LMV - PubMed Abstract:
Androcam replaces calmodulin as a tissue-specific myosin VI light chain on the actin cones that mediate D. melanogaster spermatid individualization. We show that the androcam structure and its binding to the myosin VI structural (Insert 2) and regulatory (IQ) light chain sites are distinct from those of calmodulin and provide a basis for specialized myosin VI function. The androcam N lobe noncanonically binds a single Ca(2+) and is locked in a "closed" conformation, causing androcam to contact the Insert 2 site with its C lobe only. Androcam replacing calmodulin at Insert 2 will increase myosin VI lever arm flexibility, which may favor the compact monomeric form of myosin VI that functions on the actin cones by facilitating the collapse of the C-terminal region onto the motor domain. The tethered androcam N lobe could stabilize the monomer through contacts with C-terminal portions of the motor or recruit other components to the actin cones. Androcam binds the IQ site at all calcium levels, constitutively mimicking a conformation adopted by calmodulin only at intermediate calcium levels. Thus, androcam replacing calmodulin at IQ will abolish a Ca(2+)-regulated, calmodulin-mediated myosin VI structural change. We propose that the N lobe prevents androcam from interfering with other calmodulin-mediated Ca(2+) signaling events. We discuss how gene duplication and mutations that selectively stabilize one of the many conformations available to calmodulin support the molecular evolution of structurally and functionally distinct calmodulin-like proteins.
- Department of Biochemistry and Cell Biology, Rice University, Houston, TX 77005, USA.
Organizational Affiliation: