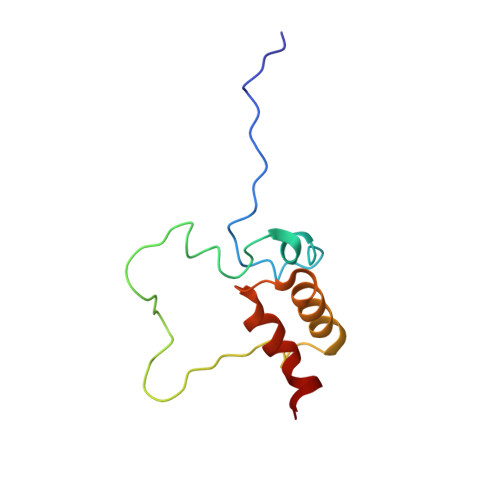
Structural Elucidation and Functional Characterization of the Hyaloperonospora arabidopsidis Effector Protein ATR13.
Leonelli, L., Pelton, J., Schoeffler, A., Dahlbeck, D., Berger, J., Wemmer, D.E., Staskawicz, B.(2011) PLoS Pathog 7: e1002428-e1002428
- PubMed: 22194684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1002428
- Primary Citation Related Structures:
2LAI - PubMed Abstract:
The oomycete Hyaloperonospora arabidopsidis (Hpa) is the causal agent of downy mildew on the model plant Arabidopsis thaliana and has been adapted as a model system to investigate pathogen virulence strategies and plant disease resistance mechanisms. Recognition of Hpa infection occurs when plant resistance proteins (R-genes) detect the presence or activity of pathogen-derived protein effectors delivered to the plant host. This study examines the Hpa effector ATR13 Emco5 and its recognition by RPP13-Nd, the cognate R-gene that triggers programmed cell death (HR) in the presence of recognized ATR13 variants. Herein, we use NMR to solve the backbone structure of ATR13 Emco5, revealing both a helical domain and a disordered internal loop. Additionally, we use site-directed and random mutagenesis to identify several amino acid residues involved in the recognition response conferred by RPP13-Nd. Using our structure as a scaffold, we map these residues to one of two surface-exposed patches of residues under diversifying selection. Exploring possible roles of the disordered region within the ATR13 structure, we perform domain swapping experiments and identify a peptide sequence involved in nucleolar localization. We conclude that ATR13 is a highly dynamic protein with no clear structural homologues that contains two surface-exposed patches of polymorphism, only one of which is involved in RPP13-Nd recognition specificity.
- Department of Plant and Microbial Biology, University of California, Berkeley, California, United States of America.
Organizational Affiliation: