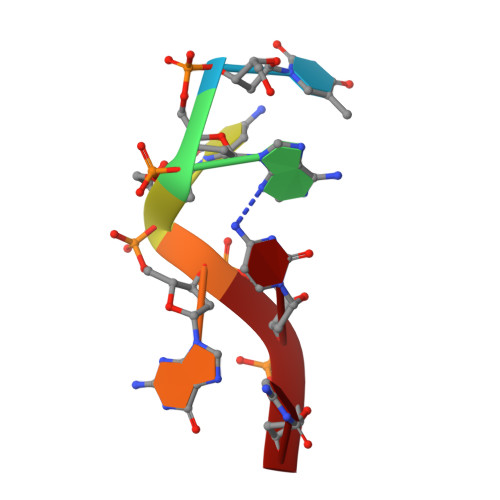
Platinated DNA affects zinc finger conformation. Interaction of a platinated single-stranded oligonucleotide and the C-terminal zinc finger of nucleocapsid protein HIVNCp7.
Quintal, S., Viegas, A., Erhardt, S., Cabrita, E.J., Farrell, N.P.(2012) Biochemistry 51: 1752-1761
- PubMed: 22303928 Search on PubMed
- DOI: https://doi.org/10.1021/bi201834g
- Primary Citation Related Structures:
2L44, 2L45, 2L46 - PubMed Abstract:
This paper describes for the first time the intimate molecular details of the association between a platinated oligonucleotide and a zinc finger peptide. Site-specific platination of the guanine in a single-stranded hexanucleotide gave {[Pt(dien)d(5'-TACGCC-3')], Pt(dien)(6-mer)} (II) characterized by mass spectrometry and (1)H nuclear magnetic resonance (NMR) spectroscopy. The work extends the study of platinum-nucleobase complex-zinc finger interactions using small molecules such as [Pt(dien)(9-EtGua)](2+) (I). The structure of the (34-52) C-terminal finger of HIV nucleocapsid protein HIVNCp7 (ZF1) was characterized by (1)H NMR spectroscopy and compared with that of the N-terminal single finger and the two-finger "intact" NCp7. Interaction of II with ZF1 results in significant changes in comparison to the "free" uncomplexed hexanucleotide; the major changes occurring for Trp37 resonances that are broadened and moved upfield, and other major shifts are for Gln45 (Hε21, Hγ3, Qβ), Met46 (NH, Hγ2), Lys47 (NH, Qγ), and Glu50 (Hγ2, Hγ3). The Zn-Cys/His chemical shifts show only marginal deviations. The solution structures of ZF1 and the 6-mer-ZF1 and II-ZF1 adducts were calculated from the nuclear Overhauser effect spectroscopy-derived distance constraints. The DNA position in the II-ZF1 adduct is completely different than in the absence of platinum. Major differences are the appearance of new Met46-Cyt6 H5 and Trp37-Cyt5 H5 contacts but severe weakening of the Trp37-Gua4 contact, attributed to the steric effects caused by Gua4 platination, accompanied by a change in the position of the aromatic ring. The results demonstrate the feasibility of targeting specific ZF motifs with DNA-tethered coordination compounds, such as Pt compounds and Co macrocycles, with implications for drug targetting and indeed the intimate mechanisms of DNA repair of platinated DNA.
- Department of Chemistry, Virginia Commonwealth University, Richmond, Virginia 23284, United States.
Organizational Affiliation: