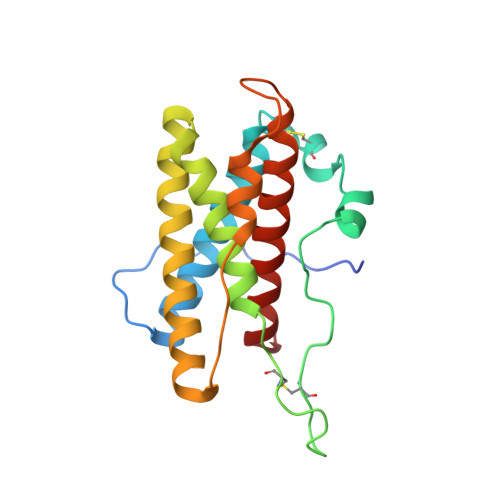
Conservation of functional sites on interleukin-6 and implications for evolution of signaling complex assembly and therapeutic intervention.
Veverka, V., Baker, T., Redpath, N.T., Carrington, B., Muskett, F.W., Taylor, R.J., Lawson, A.D., Henry, A.J., Carr, M.D.(2012) J Biological Chem 287: 40043-40050
- PubMed: 23027872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.405597
- Primary Citation Related Structures:
2L3Y - PubMed Abstract:
A number of secreted cytokines, such as interleukin-6 (IL-6), are attractive targets for the treatment of inflammatory diseases. We have determined the solution structure of mouse IL-6 to assess the functional significance of apparent differences in the receptor interaction sites (IL-6Rα and gp130) suggested by the fairly low degree of sequence similarity with human IL-6. Structure-based sequence alignment of mouse IL-6 and human IL-6 revealed surprising differences in the conservation of the two distinct gp130 binding sites (IIa and IIIa), which suggests a primacy for site III-mediated interactions in driving initial assembly of the IL-6/IL-6Rα/gp130 ternary complex. This is further supported by a series of direct binding experiments, which clearly demonstrate a high affinity IL-6/IL-6Rα-gp130 interaction via site III but only weak binding via site II. Collectively, our findings suggest a pathway for the evolution of the hexameric, IL-6/IL-6Rα/gp130 signaling complex and strategies for therapeutic targeting. We propose that the signaling complex originally involved specific interactions between IL-6 and IL-6Rα (site I) and between the D1 domain of gp130 and IL-6/IL-6Rα (site III), with the later inclusion of interactions between the D2 and D3 domains of gp130 and IL-6/IL-6Rα (site II) through serendipity. It seems likely that IL-6 signaling benefited from the evolution of a multipurpose, nonspecific protein interaction surface on gp130, now known as the cytokine binding homology region (site II contact surface), which fortuitously contributes to stabilization of the IL-6/IL-6Rα/gp130 signaling complex.
- Department of Biochemistry, University of Leicester, Henry Wellcome Building, Lancaster Road, Leicester LE1 9HN, United Kingdom.
Organizational Affiliation: