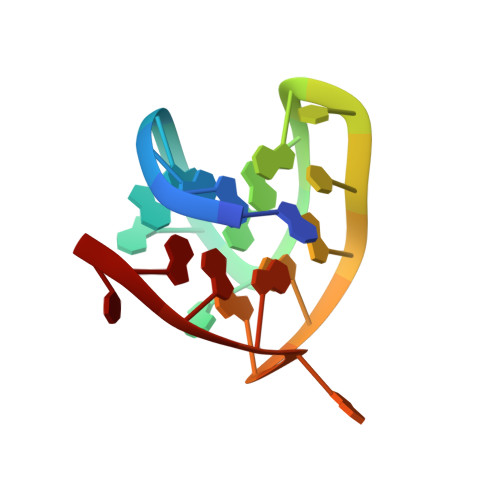
Coexistence of two distinct G-quadruplex conformations in the hTERT promoter
Lim, K.W., Lacroix, L., Yue, D.J.E., Lim, J.K.C., Lim, J.M.W., Phan, A.T.(2010) J Am Chem Soc 132: 12331-12342
- PubMed: 20704263 Search on PubMed
- DOI: https://doi.org/10.1021/ja101252n
- Primary Citation Related Structures:
2KZD, 2KZE - PubMed Abstract:
The catalytic subunit of human telomerase, hTERT, actively elongates the 3' end of the telomere in most cancer cells. The hTERT promoter, which contains many guanine-rich stretches on the same DNA strand, exhibits an exceptional potential for G-quadruplex formation. Here we show that one particular G-rich sequence in this region coexists in two G-quadruplex conformations in potassium solution: a (3 + 1) and a parallel-stranded G-quadruplexes. We present the NMR solution structures of both conformations, each comprising several robust structural elements, among which include the (3 + 1) and all-parallel G-tetrad cores, single-residue double-chain-reversal loops, and a capping A.T base pair. A combination of NMR and CD techniques, complemented with sequence modifications and variations of experimental condition, allowed us to better understand the coexistence of the two G-quadruplex conformations in equilibrium and how different structural elements conspire to favor a particular form.
- School of Physical and Mathematical Sciences, Nanyang Technological University, Singapore.
Organizational Affiliation: