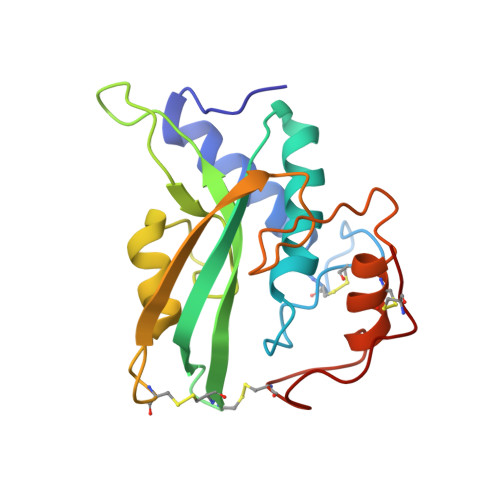
The NMR structure of protein-glutaminase from Chryseobacterium proteolyticum.
Kumeta, H., Miwa, N., Ogura, K., Kai, Y., Mizukoshi, T., Shimba, N., Suzuki, E., Inagaki, F.(2010) J Biomol NMR 46: 251-255
- PubMed: 20195702 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-010-9399-7
- Primary Citation Related Structures:
2KSV - Laboratory of Structural Biology, Graduate School of Pharmaceutical Sciences, Hokkaido University, Kita 12 Nishi 6, Kita-ku, Sapporo, 060-0812, Japan.
Organizational Affiliation: