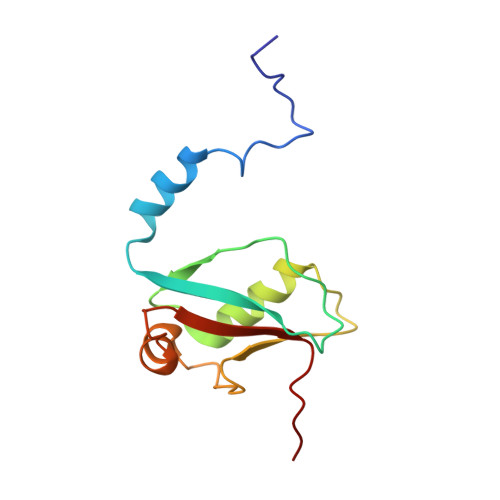
Solution structure of Atg8 reveals conformational polymorphism of the N-terminal domain
Schwarten, M., Stoldt, M., Mohrluder, J., Willbold, D.(2010) Biochem Biophys Res Commun
- PubMed: 20382112 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2010.04.043
- Primary Citation Related Structures:
2KQ7 - PubMed Abstract:
During autophagy a crescent shaped like membrane is formed, which engulfs the material that is to be degraded. This membrane grows further until its edges fuse to form the double membrane covered autophagosome. Atg8 is a protein, which is required for this initial step of autophagy. Therefore, a multistage conjugation process of newly synthesized Atg8 to phosphatidylethanolamine is of critical importance. Here we present the high resolution structure of unprocessed Atg8 determined by nuclear magnetic resonance spectroscopy. Its C-terminal subdomain shows a well-defined ubiquitin-like fold with slightly elevated mobility in the pico- to nanosecond timescale as determined by heteronuclear NOE data. In comparison to unprocessed Atg8, cleaved Atg8(G116) shows a decreased mobility behaviour. The N-terminal domain adopts different conformations within the micro- to millisecond timescale. The possible biological relevance of the differences in dynamic behaviours between both subdomains as well as between the cleaved and uncleaved forms is discussed.
- Institut für Strukturbiologie und Biophysik, ISB-3, Forschungszentrum Jülich, 52425 Jülich, Germany. m.schwarten@fz-juelich.de
Organizational Affiliation: