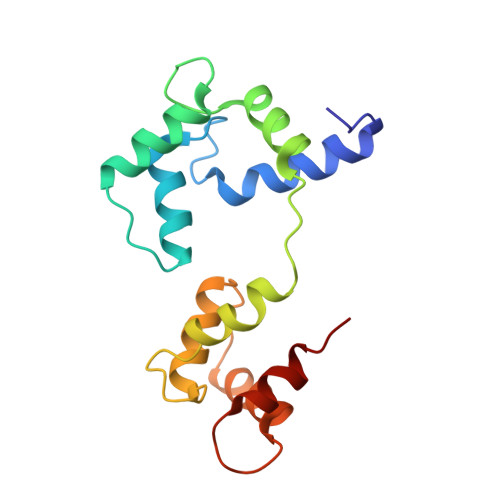
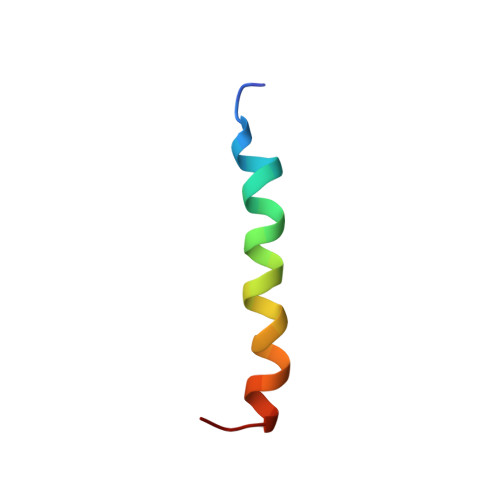
Calmodulin wraps around its binding domain in the plasma membrane Ca2+ pump anchored by a novel 18-1 motif.
Juranic, N., Atanasova, E., Filoteo, A.G., Macura, S., Prendergast, F.G., Penniston, J.T., Strehler, E.E.(2010) J Biological Chem 285: 4015-4024
- PubMed: 19996092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.060491
- Primary Citation Related Structures:
2KNE - PubMed Abstract:
Using solution NMR spectroscopy, we obtained the structure of Ca(2+)-calmodulin (holoCaM) in complex with peptide C28 from the binding domain of the plasma membrane Ca(2+)-ATPase (PMCA) pump isoform 4b. This provides the first atomic resolution insight into the binding mode of holoCaM to the full-length binding domain of PMCA. Structural comparison of the previously determined holoCaM.C20 complex with this holoCaM.C28 complex supports the idea that the initial binding step is represented by (holoCaM.C20) and the final bound complex by (holoCaM.C28). This affirms the existing multi-step kinetic model of PMCA4b activation by CaM. The complex exhibits a new binding motif in which holoCaM is wrapped around helical C28 peptide using two anchoring residues from the peptide at relative positions 18 and 1. The anchors correspond to Phe-1110 and Trp-1093, respectively, in full-length PMCA4b, and the peptide and CaM are oriented in an anti-parallel manner. This is a greater sequence distance between anchors than in any of the known holoCaM complexes with a helical peptide. Analysis of the geometry of holoCaM-peptide binding for the cases where the target peptide adopts an alpha(D)-helix with its anchors buried in the main hydrophobic pockets of the two CaM lobes establishes that only relative sequential positions of 10, 14, 17, and 18 are allowed for the second anchor.
- From the Departments of Biochemistry and Molecular Biology, Mayo Clinic College of Medicine, Rochester, Minnesota 55905. Electronic address: juranic.nenad@mayo.edu.
Organizational Affiliation: