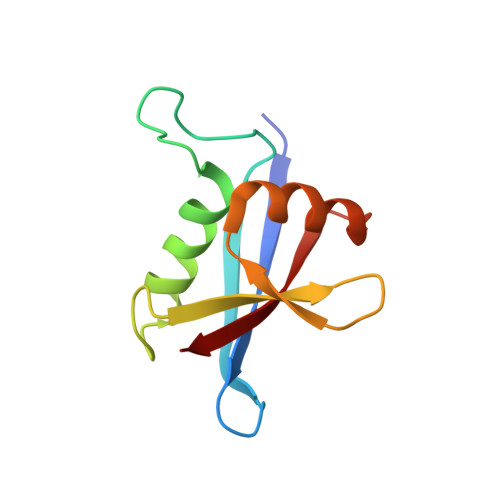
The NMR structure of the p62 PB1 domain, a key protein in autophagy and NF-kappaB signaling pathway
Saio, T., Yokochi, M., Inagaki, F.(2009) J Biomol NMR 45: 335-341
- PubMed: 19728111 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-009-9370-7
- Primary Citation Related Structures:
2KKC - Graduate School of Life Science, Hokkaido University, Sapporo 001-0021, Japan.
Organizational Affiliation: