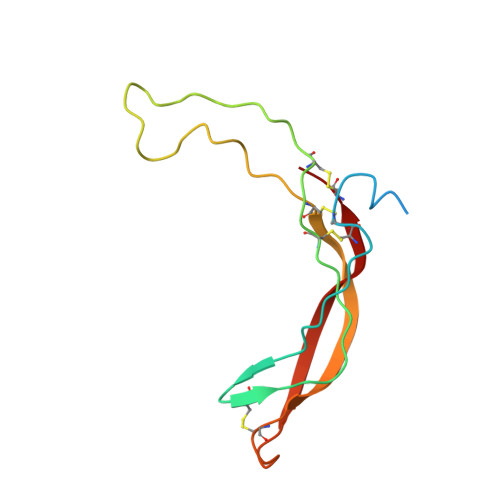
NMR structure of the Wnt modulator protein Sclerostin
Weidauer, S.E., Schmieder, P., Beerbaum, M., Schmitz, W., Oschkinat, H., Mueller, T.D.(2009) Biochem Biophys Res Commun 380: 160-165
- PubMed: 19166819 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.01.062
- Primary Citation Related Structures:
2KD3 - PubMed Abstract:
Sclerostin has been identified as a negative regulator of bone growth. Initially it was considered that Sclerostin performs its regulatory function via acting as a modulator of bone morphogenetic proteins (BMPs) similar to known examples such as Noggin, Chordin, and members of the DAN family. Recent findings, however, show that Sclerostin interferes with the Wnt signaling pathway due to binding to the Wnt co-receptor LRP5 thereby modulating bone growth. As Sclerostin is exclusively produced by osteocytes located in bones, neutralization of its bone-inhibiting functions makes it a highly interesting target for an osteoanabolic therapeutic approach in diseases characterized by bone loss, such as osteoporosis. Despite the huge interest in Sclerostin inhibitors the molecular basis of its function and its interaction with components of the Wnt signaling cascade has remained unclear. Here, we present the NMR structure of murine Sclerostin providing the first insights how Sclerostin might bind to LRP5.
- Lehrstuhl für Botanik I-Molekulare Pflanzenphysiologie und Biophysik, Julius-von-Sachs Institut für Biowissenschaften (Biozentrum) der Universität Würzburg, Julius-von-Sachs Platz 2, D-97082 Würzburg, Germany.
Organizational Affiliation: