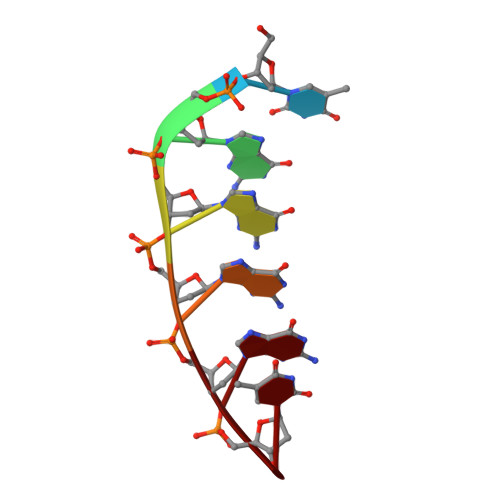
Structural and thermodynamic studies of the interaction of distamycin A with the parallel quadruplex structure [d(TGGGGT)]4
Martino, L., Virno, A., Pagano, B., Virgilio, A., Di Micco, S., Galeone, A., Giancola, C., Bifulco, G., Mayol, L., Randazzo, A.(2007) J Am Chem Soc 129: 16048-16056
- PubMed: 18052170 Search on PubMed
- DOI: https://doi.org/10.1021/ja075710k
- Primary Citation Related Structures:
2JT7 - PubMed Abstract:
The complex between distamycin A and the parallel DNA quadruplex [d(TGGGGT)]4 has been studied by 1H NMR spectroscopy and isothermal titration calorimetry (ITC). To unambiguously assert that distamycin A interacts with the grooves of the quadruplex [d(TGGGGT)]4, we have analyzed the NMR titration profile of a modified quadruplex, namely [d(TGGMeGGT)]4, and we have applied the recently developed differential frequency-saturation transfer difference (DF-STD) method, for assessing the ligand-DNA binding mode. The three-dimensional structure of the 4:1 distamycin A/[d(TGGGGT)]4 complex has been determined by an in-depth NMR study followed by dynamics and mechanics calculations. All results unequivocally indicate that distamycin molecules interact with [d(TGGGGT)]4 in a 4:1 binding mode, with two antiparallel distamycin dimers that bind simultaneously two opposite grooves of the quadruplex. The affinity between distamycin A and [d(TGGGGT)]4 enhances ( approximately 10-fold) when the ratio of distamycin A to the quadruplex is increased. In this paper we report the first three-dimensional structure of a groove-binder molecule complexed to a DNA quadruplex structure.
- Dipartimento di Chimica P. Corradini, Università degli Studi di Napoli Federico II, via Cintia, I-80126, Napoli, Italy.
Organizational Affiliation: