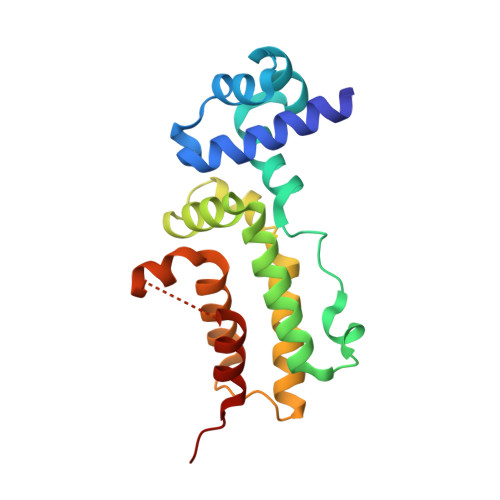
Structural Investigation of Transcriptional Regulator Hlyiir: Influence of a Disordered Region on Protein Fold and Dimerization.
Kovalevskiy, O.V., Solonin, A.S., Antson, A.A.(2010) Proteins 78: 1870
- PubMed: 20225260 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22700
- Primary Citation Related Structures:
2JK3, 2WV1 - PubMed Abstract:
B. cereus HlyIIR belongs to the TetR family of dimeric transcriptional regulators. Unlike other members of the TetR family, HlyIIR contains an insert between alpha-helices alpha8 and alpha9, which is located at the subunit-subunit interface. N-terminal segment of this insert (amino acids, Pro161-Ser169) forms a short alpha-helix alpha8* that occupies a complementary cavity on the surface of the adjacent subunit, whereas the C-terminal segment comprising 16 amino acids (Leu170-Glu185) is disordered. To understand whether this disordered segment is important for protein's function, we determined crystal structures of two engineered HlyIIR proteins where this segment was either substituted by a seven-residue flexible Ser-Gly linker or replaced by a cleavable peptide containing proteolytic sites at both ends. Unexpectedly, alteration or proteolytic removal of the disordered segment resulted in changes in protein's conformation and in a remarkable rearrangement at the subunit-subunit interface. X-ray structures of the two engineered proteins revealed an unusual plasticity at the dimerization interface of HlyIIR enabling it to form dimers stabilized by different sets of interactions. Structural comparison indicates that in spite of the flexible nature of the disordered segment, it is critical for maintaining the native structure as it influences the position of alpha8*. The data demonstrate how disordered loops on protein surfaces may affect folding and subunit-subunit interactions.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York, United Kingdom. oleg@ysbl.york.ac.uk
Organizational Affiliation: