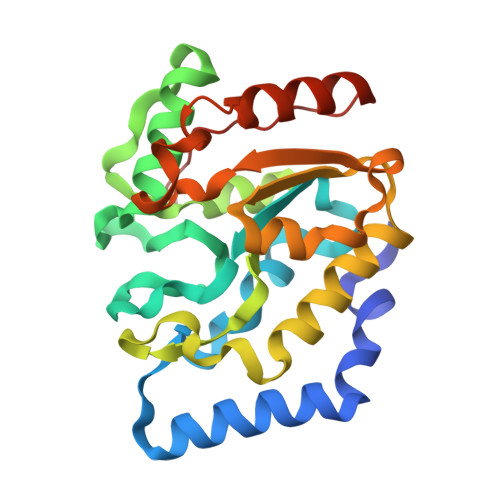
Structure of Uracil-DNA N-Glycosylase (Ung) from Vibrio Cholerae. Mapping Temperature Adaptation Through Structural and Mutational Analysis.
Raeder, I.L.U., Moe, E., Willassen, N.P., Smalas, A.O., Leiros, I.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 130
- PubMed: 20124707 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109052063
- Primary Citation Related Structures:
2JHQ - PubMed Abstract:
The crystal structure of Vibrio cholerae uracil-DNA N-glycosylase (vcUNG) has been determined to 1.5 A resolution. Based on this structure, a homology model of Aliivibrio salmonicida uracil-DNA N-glycosylase (asUNG) was built. A previous study demonstrated that asUNG possesses typical cold-adapted features compared with vcUNG, such as a higher catalytic efficiency owing to increased substrate affinity. Specific amino-acid substitutions in asUNG were suggested to be responsible for the increased substrate affinity and the elevated catalytic efficiency by increasing the positive surface charge in the DNA-binding region. The temperature adaptation of these enzymes has been investigated using structural and mutational analyses, in which mutations of vcUNG demonstrated an increased substrate affinity that more resembled that of asUNG. Visualization of surface potentials revealed a more positive potential for asUNG compared with vcUNG; a modelled double mutant of vcUNG had a potential around the substrate-binding region that was more like that of asUNG, thus rationalizing the results obtained from the kinetic studies.
- Department of Chemistry, Faculty of Science, University of Tromsø, N-9037 Tromsø, Norway.
Organizational Affiliation: