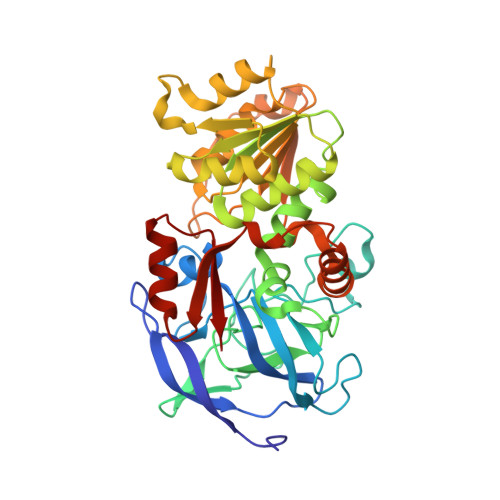
Structural Evidence for a Ligand Coordination Switch in Liver Alcohol Dehydrogenase
Meijers, R., Adolph, H.W., Dauter, Z., Wilson, K.S., Lamzin, V.S., Cedergren-Zeppezauer, E.S.(2007) Biochemistry 46: 5446
- PubMed: 17429946 Search on PubMed
- DOI: https://doi.org/10.1021/bi6023594
- Primary Citation Related Structures:
2JHF, 2JHG - PubMed Abstract:
The use of substrate analogues as inhibitors provides a way to understand and manipulate enzyme function. Here we report two 1 A resolution crystal structures of liver alcohol dehydrogenase in complex with NADH and two inhibitors: dimethyl sulfoxide and isobutyramide. Both structures present a dynamic state of inhibition. In the dimethyl sulfoxide complex structure, the inhibitor is caught in transition on its way to the active site using a flash-freezing protocol and a cadmium-substituted enzyme. One inhibitor molecule is partly located in the first and partly in the second coordination sphere of the active site metal. A hydroxide ion bound to the active site metal lies close to the pyridine ring of NADH, which is puckered in a twisted boat conformation. The cadmium ion is coordinated by both the hydroxide ion and the inhibitor molecule, providing structural evidence of a coordination switch at the active site metal ion. The structure of the isobutyramide complex reveals the partial formation of an adduct between the isobutyramide inhibitor and NADH. It provides evidence of the contribution of a shift from the keto to the enol tautomer during aldehyde reduction. The different positions of the inhibitors further refine the knowledge of the dynamics of the enzyme mechanism and explain how the crowded active site can facilitate the presence of a substrate and a metal-bound hydroxide ion.
- European Molecular Biology Laboratory, Hamburg Outstation, c/o DESY, Notkestrasse 85, D-22603 Hamburg, Federal Republic of Germany. rob.meijers@synchrotron-soleil.fr
Organizational Affiliation: