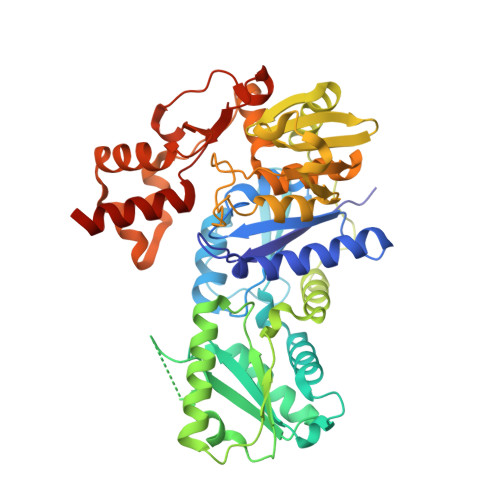
The Crystal Structure of Dr2241 from Deinococcus Radiodurans at 1.9 A Resolution Reveals a Multi-Domain Protein with Structural Similarity to Chelatases But Also with Two Additional Novel Domains
Leiros, H.-K.S., Mcsweeney, S.M.(2007) J Struct Biol 159: 92
- PubMed: 17448684 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2007.02.009
- Primary Citation Related Structures:
2JH3 - PubMed Abstract:
A unique family of proteins have been identified in the Deinococcus genus with an N-terminal cobalamin (vitamin B(12)) chelatase domain denoted CbiX and an additional unique C-terminal domain with unknown function. Here we report the first crystal structure from this new family of proteins with the structure of Deinococcus radiodurans protein DR2241. The structure reveals a multi-domain protein where domains A (residues 1-132) has the same fold as the small CbiX (CbiX(S)), domains A and B (residues 1-272) follow the chelatase super-family fold and the two additional unique domains C and D have no structural homologues. Domain D harbours the sequence motifs CxxC and CxxxC, in which DR2241 gives the first evidence that these motifs bind a [4Fe-4S] iron-sulphur cluster. In solution there are indications of multimeric forms, and in the crystallographic asymmetric unit a tetramer is found where domains C and D are involved in stabilising the tetrameric assembly.
- Macromolecular Crystallography Group, European Synchrotron Radiation Facility (ESRF), BP 220, 6, Rue Jules Horowitz, F-38043 Grenoble Cedex 09, France.
Organizational Affiliation: