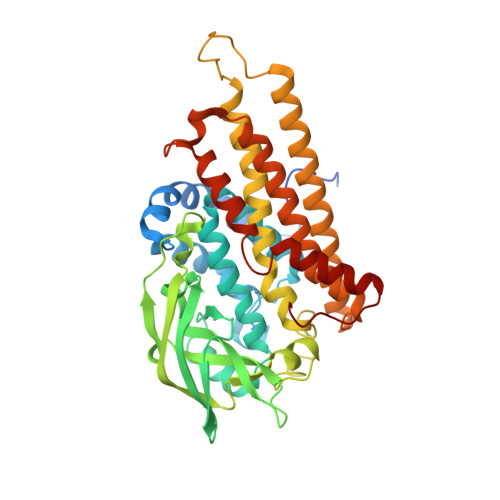
Structure of the Monooxygenase Component of a Two-Component Flavoprotein Monooxygenase.
Alfieri, A., Fersini, F., Ruangchan, N., Prongjit, M., Chaiyen, P., Mattevi, A.(2007) Proc Natl Acad Sci U S A 104: 1177
- PubMed: 17227849 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0608381104
- Primary Citation Related Structures:
2JBR, 2JBS, 2JBT - PubMed Abstract:
p-Hydroxyphenylacetate hydroxylase from Acinetobacter baumannii is a two-component system consisting of a NADH-dependent FMN reductase and a monooxygenase (C2) that uses reduced FMN as substrate. The crystal structures of C2 in the ligand-free and substrate-bound forms reveal a preorganized pocket that binds reduced FMN without large conformational changes. The Phe-266 side chain swings out to provide the space for binding p-hydroxyphenylacetate that is oriented orthogonal to the flavin ring. The geometry of the substrate-binding site of C2 is significantly different from that of p-hydroxybenzoate hydroxylase, a single-component flavoenzyme that catalyzes a similar reaction. The C2 overall structure resembles the folding of medium-chain acyl-CoA dehydrogenase. An outstanding feature in the C2 structure is a cavity located in front of reduced FMN; it has a spherical shape with a 1.9-A radius and a 29-A3 volume and is interposed between the flavin C4a atom and the substrate atom to be hydroxylated. The shape and position of this cavity are perfectly fit for housing the oxygen atoms of the flavin C4a-hydroperoxide intermediate that is formed upon reaction of the C2-bound reduced flavin with molecular oxygen. The side chain of His-396 is predicted to act as a hydrogen-bond donor to the oxygen atoms of the intermediate. This architecture promotes the nucleophilic attack of the substrate onto the terminal oxygen of the hydroperoxyflavin. Comparative analysis with the structures of other flavoenzymes indicates that a distinctive feature of monooxygenases is the presence of specific cavities that encapsulate and stabilize the crucial hydroperoxyflavin intermediate.
- Department of Genetics and Microbiology, University of Pavia, Via Ferrata 1, 27100 Pavia, Italy.
Organizational Affiliation: