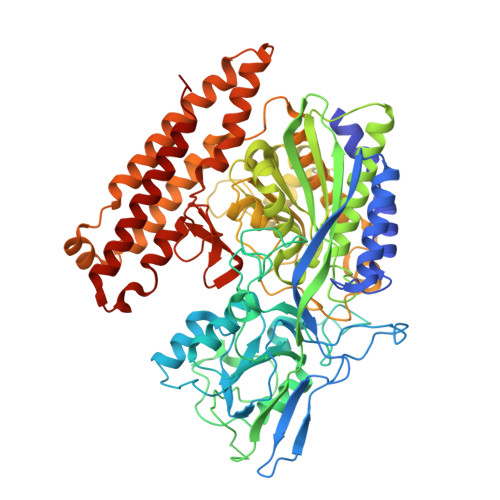
Human Glutamate Carboxypeptidase II Inhibition: Structures of Gcpii in Complex with Two Potent Inhibitors, Quisqualate and 2-Pmpa.
Mesters, J.R., Henning, K., Hilgenfeld, R.(2007) Acta Crystallogr D Biol Crystallogr 63: 508
- PubMed: 17372356 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490700902X
- Primary Citation Related Structures:
2JBJ, 2JBK - PubMed Abstract:
Human glutamate carboxypeptidase II (GCPII) occurs in the central nervous system as well as in human prostate (where it is called prostate-specific membrane antigen; PSMA). Inhibitors of the enzyme have been shown to provide neuroprotection, but may also be useful for the detection, imaging and treatment of prostate cancer. Crystal structures were determined of the extracellular part of GCPII (amino-acid residues 44-750) in complex with two potent inhibitors, quisqualate and 2-PMPA (the strongest GCPII inhibitor to date), at resolutions of 3.0 and 2.2 A, respectively. In addition, models were constructed for binding of the inhibitors willardiine, homoibotenate, L-2-amino-4-phosphonobutanoic acid and L-serine-O-sulfate to the S1' site of the enzyme. The common denominator for high-affinity binding to the S1' site is the formation of two strong salt bridges.
- Institute of Biochemistry, Center for Structural and Cell Biology In Medicine, University of Lübeck, Ratzeburger Allee 160, Lübeck 23538, Germany. mesters@biochem.uni-luebeck.de
Organizational Affiliation: