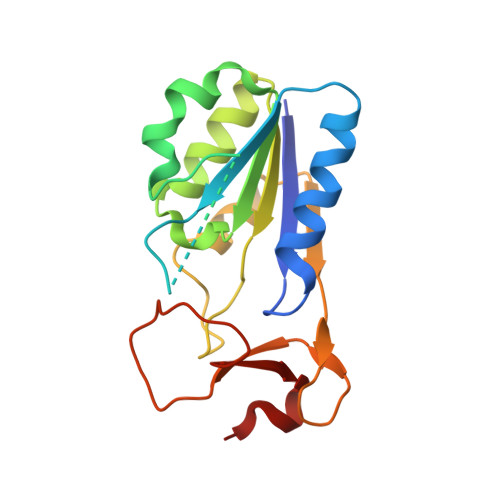
Structure of Vaccinia Virus Thymidine Kinase in Complex with Dttp: Insights for Drug Design.
El Omari, K., Solaroli, N., Karlsson, A., Balzarini, J., Stammers, D.K.(2006) BMC Struct Biol 6: 22
- PubMed: 17062140 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-6-22
- Primary Citation Related Structures:
2J87 - PubMed Abstract:
Development of countermeasures to bioterrorist threats such as those posed by the smallpox virus (variola), include vaccination and drug development. Selective activation of nucleoside analogues by virus-encoded thymidine (dThd) kinases (TK) represents one of the most successful strategies for antiviral chemotherapy as demonstrated for anti-herpes drugs. Vaccinia virus TK is a close orthologue of variola TK but also shares a relatively high sequence identity to human type 2 TK (hTK), thus achieving drug selectivity relative to the host enzyme is challenging. In order to identify any differences compared to hTK that may be exploitable in drug design, we have determined the crystal structure of VVTK, in complex with thymidine 5'-triphosphate (dTTP). Although most of the active site residues are conserved between hTK and VVTK, we observe a difference in conformation of residues Asp-43 and Arg-45. The equivalent residues in hTK hydrogen bond to dTTP, whereas in subunit D of VVTK, Asp-43 and Arg-45 adopt a different conformation preventing interaction with this nucleotide. Asp-43 and Arg-45 are present in a flexible loop, which is disordered in subunits A, B and C. The observed difference in conformation and flexibility may also explain the ability of VVTK to phosphorylate (South)-methanocarbathymine whereas, in contrast, no substrate activity with hTK is reported for this compound. The difference in conformation for Asp-43 and Arg-45 could thus be used in drug design to generate VVTK/Variola TK-selective nucleoside analogue substrates and/or inhibitors that have lower affinity for hTK.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford OX3 7BN, UK. kamel@strubi.ox.ac.uk <kamel@strubi.ox.ac.uk>
Organizational Affiliation: