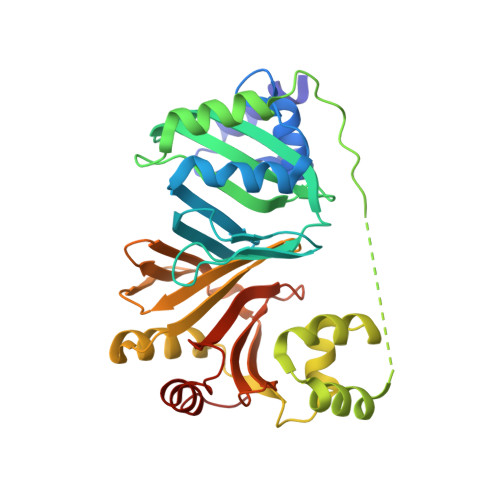
An Induced Fit Conformational Change Underlies the Binding Mechanism of the Heme Transport Proteobacteria-Protein Hems.
Schneider, S., Sharp, K.H., Barker, P.D., Paoli, M.(2006) J Biological Chem 281: 32606
- PubMed: 16943192 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M607516200
- Primary Citation Related Structures:
2J0P, 2J0R - PubMed Abstract:
Bacteria rely on their environment and/or host to acquire iron and have evolved specialized systems to sequester and transport heme. The heme uptake system HemRSTUV is common to proteobacteria, and a major challenge is to understand the molecular mechanism of heme binding and transfer between the protein molecules that underlie this heme transport relay process. In the Gram-negative pathogen Yersinia enterocolitica, the HemRSTUV system culminates with the cytoplasmic recipient HemS, which stores and delivers heme for cellular needs. HemS belongs to a family of proteins essential and unique to proteobacteria. Here we report on the binding mechanism of HemS based on structural data from its apo- and ligand-loaded forms. This heme carrier protein associates with its cargo through a novel, partly preformed binding pocket, formed between a large beta-sheet dome and a three-helix subdomain. In addition to a histidine interacting with the iron, the complex is stabilized by a distal non-coordinating arginine that packs along the porphyrin plane and extensive electrostatic contacts that firmly anchor the heme propionate groups within the protein. Comparison of apo- and ligand-bound HemS crystal structures reveals striking conformational changes that underlie a "heme-induced fit" binding mechanism. Local shifts in amino acid positions combine with global, rigid body-like domain movements, and together, these bring about a switch from an open, apo-form to a closed, bound state. This is the first report in which both liganded and unliganded forms of a heme transport protein are described, thus providing penetrating insights into its mechanism of heme binding and release.
- School of Pharmacy and Centre for Biomolecular Sciences, University of Nottingham, University Park, Nottingham NG7 2RD.
Organizational Affiliation: