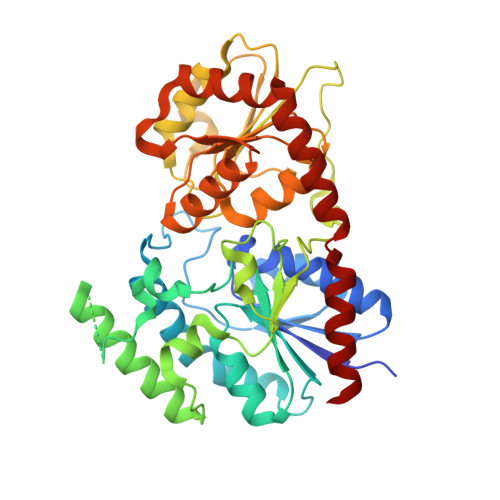
The Crystal Structure of Two Macrolide Glycosyltransferases Provides a Blueprint for Host Cell Antibiotic Immunity.
Bolam, D.N., Roberts, S.M., Proctor, M.R., Turkenburg, J.P., Dodson, E.J., Martinez-Fleites, C., Yang, M., Davis, B.G., Davies, G.J., Gilbert, H.J.(2007) Proc Natl Acad Sci U S A 104: 5336
- PubMed: 17376874 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0607897104
- Primary Citation Related Structures:
2IYA, 2IYF - PubMed Abstract:
Glycosylation of macrolide antibiotics confers host cell immunity from endogenous and exogenous agents. The Streptomyces antibioticus glycosyltransferases, OleI and OleD, glycosylate and inactivate oleandomycin and diverse macrolides including erythromycin, respectively. The structure of these enzyme-ligand complexes, in tandem with kinetic analysis of site-directed variants, provide insight into the interaction of macrolides with their synthetic apparatus. Erythromycin binds to OleD and the 23S RNA of its target ribosome in the same conformation and, although the antibiotic contains a large number of polar groups, its interaction with these macromolecules is primarily through hydrophobic contacts. Erythromycin and oleandomycin, when bound to OleD and OleI, respectively, adopt different conformations, reflecting a subtle effect on sugar positioning by virtue of a single change in the macrolide backbone. The data reported here provide structural insight into the mechanism of resistance to both endogenous and exogenous antibiotics, and will provide a platform for the future redesign of these catalysts for antibiotic remodelling.
- Institute for Cell and Molecular Biosciences, The Medical School, University of Newcastle upon Tyne, Framlington Place, Newcastle upon Tyne NE2 4HH, UK.
Organizational Affiliation: