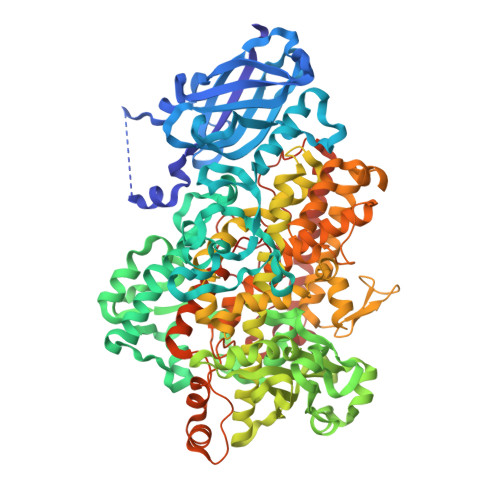
Crystal Structures of Vegetative Soybean Lipoxygenase Vlx-B and Vlx-D, and Comparisons with Seed Isoforms Lox-1 and Lox-3.
Youn, B., Sellhorn, G.E., Mirchel, R.J., Gaffney, B.J., Grimes, H.D., Kang, C.(2006) Proteins 65: 1008
- PubMed: 17022084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.21182
- Primary Citation Related Structures:
2IUJ, 2IUK - PubMed Abstract:
The lipoxygenase family of lipid-peroxidizing, nonheme iron dioxygenases form products that are precursors for diverse physiological processes in both plants and animals. In soybean (Glycine max), five vegetative isoforms, VLX-A, VLX-B, VLX-C, VLX-D, VLX-E, and four seed isoforms LOX-1, LOX-2, LOX-3a, LOX-3b have been identified. In this study, we determined the crystal structures of the substrate-free forms of two major vegetative isoforms, with distinct enzymatic characteristics, VLX-B and VLX-D. Their structures are similar to the two seed isoforms, LOX-1 and LOX-3, having two domains with similar secondary structural elements: a beta-barrel N-terminal domain containing highly flexible loops and an alpha-helix-rich C-terminal catalytic domain. Detailed comparison of the structures of these two vegetative isoforms with the structures of LOX-1 and LOX-3 reveals important differences that help explain distinct aspects of the activity and positional specificity of these enzymes. In particular, the shape of the three branches of the internal subcavity, corresponding to substrate-binding and O(2) access, differs among the isoforms in a manner that reflects the differences in positional specificities.
- School of Molecular Biosciences, Washington State University, Pullman, Washington 99164-4660, USA.
Organizational Affiliation: