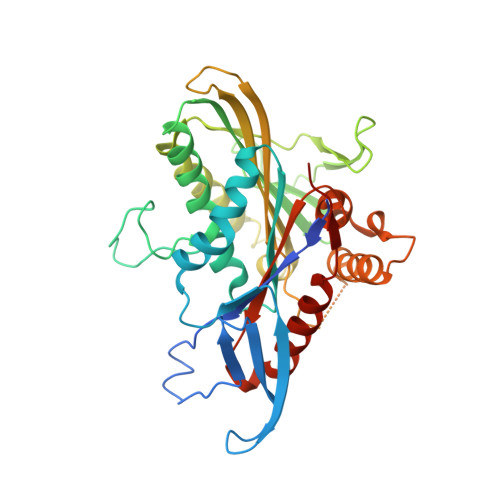
Structure of human Eg5 in complex with a new monastrol-based inhibitor bound in the R configuration.
Garcia-Saez, I., DeBonis, S., Lopez, R., Trucco, F., Rousseau, B., Thuery, P., Kozielski, F.(2007) J Biol Chem 282: 9740-9747
- PubMed: 17251189
- DOI: https://doi.org/10.1074/jbc.M608883200
- Primary Citation of Related Structures:
2IEH - PubMed Abstract:
Drugs that target mitotic spindle proteins have been proven useful for tackling tumor growth. Eg5, a kinesin-5 family member, represents a potential target, since its inhibition leads to prolonged mitotic arrest through the activation of the mitotic checkpoint and apoptotic cell death. Monastrol, a specific dihydropyrimidine inhibitor of Eg5, shows stereo-specificity, since predominantly the (S)-, but not the (R)-, enantiomer has been shown to be the biologically active compound in vitro and in cell-based assays. Here, we solved the crystal structure (2.7A) of the complex between human Eg5 and a new keto derivative of monastrol (named mon-97), a potent antimitotic inhibitor. Surprisingly, we identified the (R)-enantiomer bound in the active site, and not, as for monastrol, the (S)-enantiomer. The absolute configuration of this more active (R)-enantiomer has been unambiguously determined via chemical correlation and x-ray analysis. Unexpectedly, both the R- and the S-forms inhibit Eg5 ATPase activity with IC(50) values of 110 and 520 nM (basal assays) and 150 nm and 650 nm (microtubule-stimulated assays), respectively. However, the difference was large enough for the protein to select the (R)- over the (S)-enantiomer. Taken together, these results show that in this new monastrol family, both (R)- and (S)-enantiomers can be active as Eg5 inhibitors. This considerably broadens the alternatives for rational drug design.
Organizational Affiliation:
Laboratoire des Moteurs Moléculaires, IBS, Institut de Biologie Structurale Jean-Pierre Ebel, CNRS-Commissariat à l'Energie Atomique (CEA)-Université Joseph Fourier, 41 rue Jules Horowitz, F-38027 Grenoble.