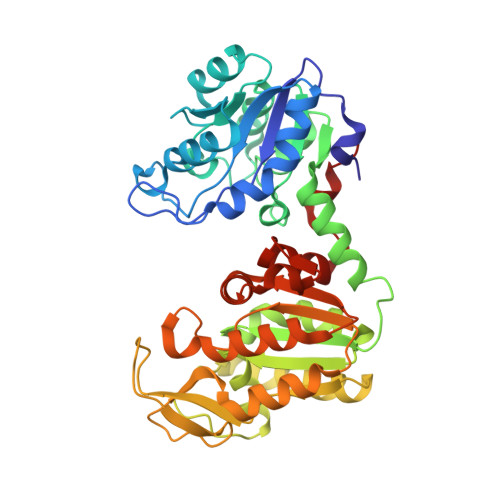
Crystal structure of Thermus caldophilus phosphoglycerate kinase in the open conformation
Lee, J.H., Im, Y.J., Bae, J., Kim, D., Kim, M.K., Kang, G.B., Lee, D.S., Eom, S.H.(2006) Biochem Biophys Res Commun 350: 1044-1049
- PubMed: 17045964 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2006.09.151
- Primary Citation Related Structures:
2IE8 - PubMed Abstract:
Phosphoglycerate kinase (PGK) is a key glycolytic enzyme that catalyzes the reversible transfer of a phosphate from 1,3-bisphosphoglycerate to ADP to form 3-phosphoglycerate and ATP in the presence of magnesium. During catalysis, a conformational change occurs that brings the N- and C-domains of PGK closer together. Here we present the 1.8A crystal structure of unliganded PGK from Thermus caldophilus (Tca). Comparison of the structure of TcaPGK (open conformation) with that of Thermotoga maritima (Tma) PGK (closed conformation) revealed that the conformational change reflects a change in the interaction between the domains. We identified Arg148 as a key residue involved in open-to-closed transition. The open conformation of TcaPGK is stabilized by an interdomain salt bridge between Arg148 and Glu375. The binding of 3-PG (or maybe 1,3-BPG) disrupts this salt bridge and, in ternary complex, the formation of new salt bridge between Arg60 and Asp197 stabilizes the closed conformation.
- Department of Life Science, Gwangju Institute of Science and Technology, Gwangju 500-712, Republic of Korea.
Organizational Affiliation: