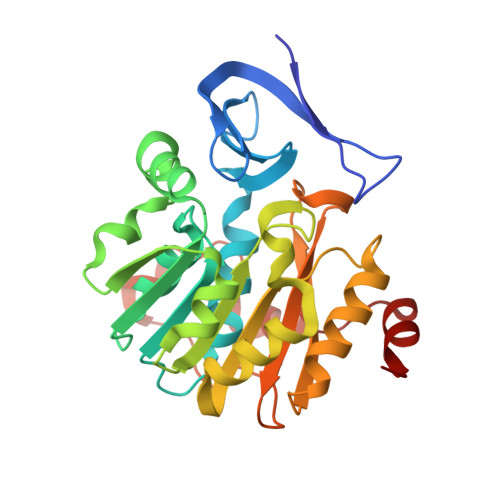
Crystal structure of Plasmodium falciparum spermidine synthase in complex with the substrate decarboxylated S-adenosylmethionine and the potent inhibitors 4MCHA and AdoDATO.
Dufe, V.T., Qiu, W., Muller, I.B., Hui, R., Walter, R.D., Al-Karadaghi, S.(2007) J Mol Biology 373: 167-177
- PubMed: 17822713 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.07.053
- Primary Citation Related Structures:
2I7C, 2PSS, 2PT6, 2PT9 - PubMed Abstract:
Plasmodium falciparum is the causative agent of the most severe type of malaria, a life-threatening disease affecting the lives of over three billion people. Factors like widespread resistance against available drugs and absence of an effective vaccine are seriously compounding control of the malaria parasite. Thus, there is an urgent need for the identification and validation of new drug targets. The enzymes of the polyamine biosynthesis pathway have been suggested as possible targets for the treatment of malaria. One of these enzymes is spermidine synthase (SPDS, putrescine aminopropyltransferase), which catalyzes the transfer of an aminopropyl moiety from decarboxylated S-adenosylmethionine (dcAdoMet) to putrescine, leading to the formation of spermidine and 5'-methylthioadenosine. Here we present the three-dimensional structure of P. falciparum spermidine synthase (pfSPDS) in apo form, in complex with dcAdoMet and two inhibitors, S-adenosyl-1,8-diamino-3-thio-octane (AdoDATO) and trans-4-methylcyclohexylamine (4MCHA). The results show that binding of dcAdoMet to pfSPDS stabilizes the conformation of the flexible gatekeeper loop of the enzyme and affects the conformation of the active-site amino acid residues, preparing the protein for binding of the second substrate. The complexes of AdoDATO and 4MCHA with pfSPDS reveal the mode of interactions of these compounds with the enzyme. While AdoDATO essentially fills the entire active-site pocket, 4MCHA only occupies part of it, which suggests that simple modifications of this compound may yield more potent inhibitors of pfSPDS.
- Department of Molecular Biophysics, Center for Molecular Protein Science, Lund University, S-221 00 Lund, Sweden.
Organizational Affiliation: