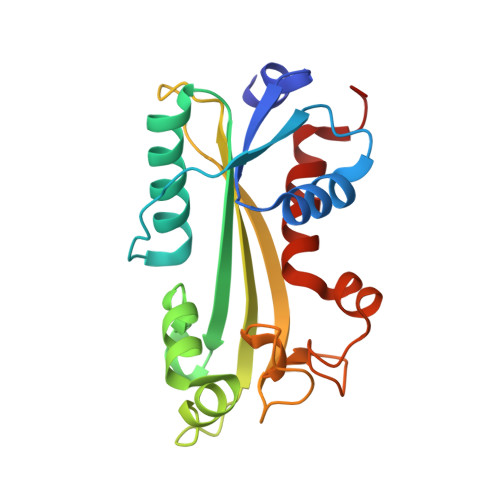
Structure of the orthorhombic form of human inosine triphosphate pyrophosphatase.
Porta, J., Kolar, C., Kozmin, S.G., Pavlov, Y.I., Borgstahl, G.E.O.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 1076-1081
- PubMed: 17077483 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309106041790
- Primary Citation Related Structures:
2I5D - PubMed Abstract:
The structure of human inosine triphosphate pyrophosphohydrolase (ITPA) has been determined using diffraction data to 1.6 A resolution. ITPA contributes to the accurate replication of DNA by cleansing cellular dNTP pools of mutagenic nucleotide purine analogs such as dITP or dXTP. A similar high-resolution unpublished structure has been deposited in the Protein Data Bank from a monoclinic and pseudo-merohedrally twinned crystal. Here, cocrystallization of ITPA with a molar ratio of XTP appears to have improved the crystals by eliminating twinning and resulted in an orthorhombic space group. However, there was no evidence for bound XTP in the structure. Comparison with substrate-bound NTPase from a thermophilic organism predicts the movement of residues within helix alpha1, the loop before alpha6 and helix alpha7 to cap off the active site when substrate is bound.
- The Eppley Institute for Research in Cancer and Allied Diseases, 987696 Nebraska Medical Center, Omaha, NE 68198-7696, USA.
Organizational Affiliation: